Laure Wynants
topic group 6 of the STRATOS initiative
Challenges and recommendations for Electronic Health Records data extraction and preparation for dynamic prediction modelling in hospitalized patients -- a practical guide
Jan 17, 2025Abstract:Dynamic predictive modeling using electronic health record (EHR) data has gained significant attention in recent years. The reliability and trustworthiness of such models depend heavily on the quality of the underlying data, which is largely determined by the stages preceding the model development: data extraction from EHR systems and data preparation. We list over forty challenges encountered during these stages and provide actionable recommendations for addressing them. These challenges are organized into four categories: cohort definition, outcome definition, feature engineering, and data cleaning. This list is designed to serve as a practical guide for data extraction engineers and researchers, supporting better practices and improving the quality and real-world applicability of dynamic prediction models in clinical settings.
Performance evaluation of predictive AI models to support medical decisions: Overview and guidance
Dec 13, 2024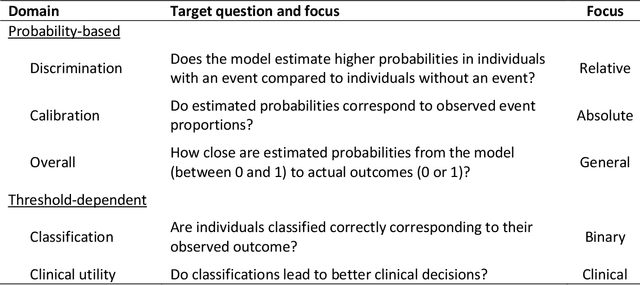
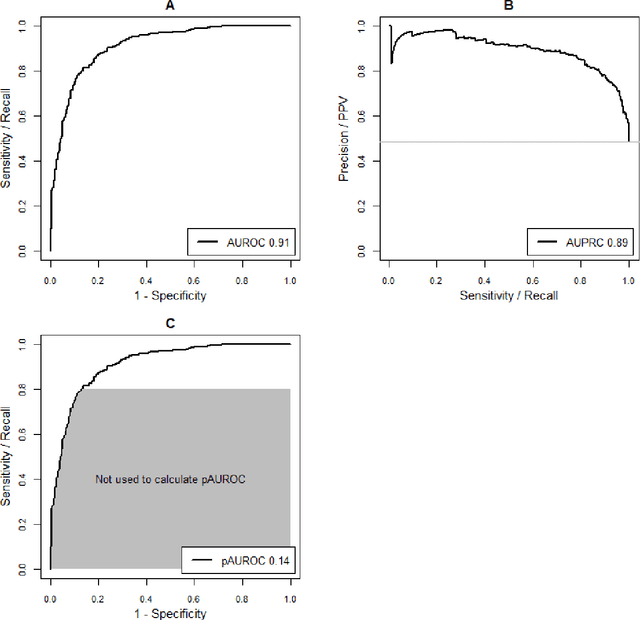
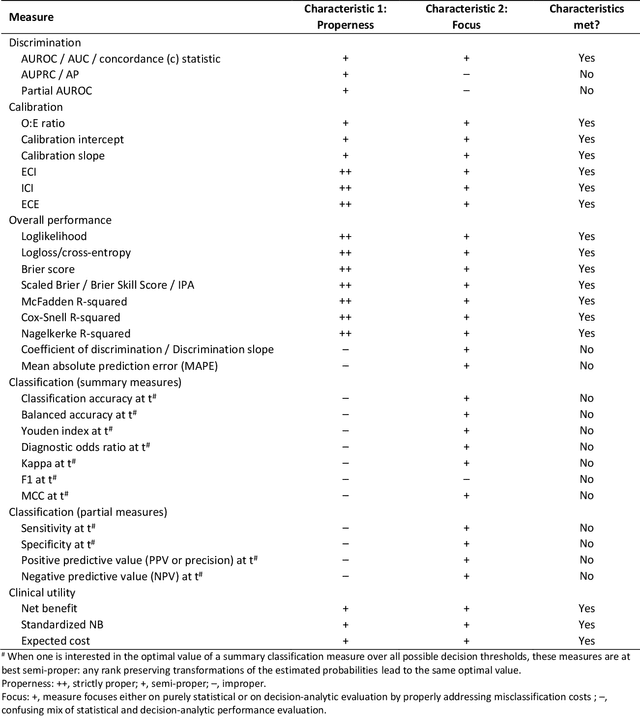
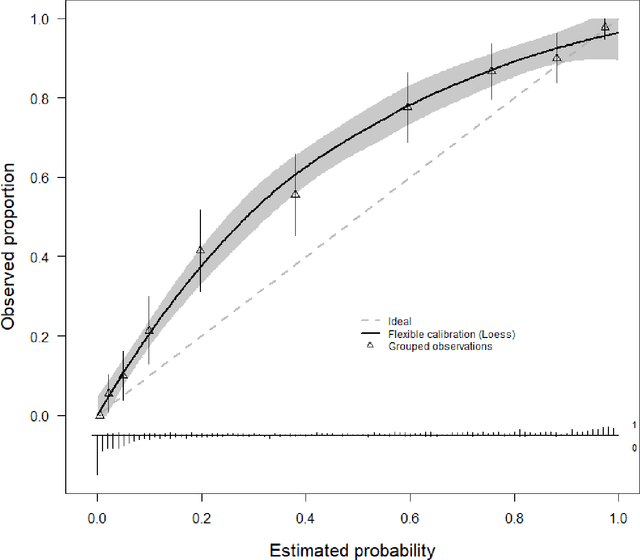
Abstract:A myriad of measures to illustrate performance of predictive artificial intelligence (AI) models have been proposed in the literature. Selecting appropriate performance measures is essential for predictive AI models that are developed to be used in medical practice, because poorly performing models may harm patients and lead to increased costs. We aim to assess the merits of classic and contemporary performance measures when validating predictive AI models for use in medical practice. We focus on models with a binary outcome. We discuss 32 performance measures covering five performance domains (discrimination, calibration, overall, classification, and clinical utility) along with accompanying graphical assessments. The first four domains cover statistical performance, the fifth domain covers decision-analytic performance. We explain why two key characteristics are important when selecting which performance measures to assess: (1) whether the measure's expected value is optimized when it is calculated using the correct probabilities (i.e., a "proper" measure), and (2) whether they reflect either purely statistical performance or decision-analytic performance by properly considering misclassification costs. Seventeen measures exhibit both characteristics, fourteen measures exhibited one characteristic, and one measure possessed neither characteristic (the F1 measure). All classification measures (such as classification accuracy and F1) are improper for clinically relevant decision thresholds other than 0.5 or the prevalence. We recommend the following measures and plots as essential to report: AUROC, calibration plot, a clinical utility measure such as net benefit with decision curve analysis, and a plot with probability distributions per outcome category.
missForestPredict -- Missing data imputation for prediction settings
Jul 02, 2024



Abstract:Prediction models are used to predict an outcome based on input variables. Missing data in input variables often occurs at model development and at prediction time. The missForestPredict R package proposes an adaptation of the missForest imputation algorithm that is fast, user-friendly and tailored for prediction settings. The algorithm iteratively imputes variables using random forests until a convergence criterion (unified for continuous and categorical variables and based on the out-of-bag error) is met. The imputation models are saved for each variable and iteration and can be applied later to new observations at prediction time. The missForestPredict package offers extended error monitoring, control over variables used in the imputation and custom initialization. This allows users to tailor the imputation to their specific needs. The missForestPredict algorithm is compared to mean/mode imputation, linear regression imputation, mice, k-nearest neighbours, bagging, miceRanger and IterativeImputer on eight simulated datasets with simulated missingness (48 scenarios) and eight large public datasets using different prediction models. missForestPredict provides competitive results in prediction settings within short computation times.
Comparison of static and dynamic random forests models for EHR data in the presence of competing risks: predicting central line-associated bloodstream infection
Apr 24, 2024



Abstract:Prognostic outcomes related to hospital admissions typically do not suffer from censoring, and can be modeled either categorically or as time-to-event. Competing events are common but often ignored. We compared the performance of random forest (RF) models to predict the risk of central line-associated bloodstream infections (CLABSI) using different outcome operationalizations. We included data from 27478 admissions to the University Hospitals Leuven, covering 30862 catheter episodes (970 CLABSI, 1466 deaths and 28426 discharges) to build static and dynamic RF models for binary (CLABSI vs no CLABSI), multinomial (CLABSI, discharge, death or no event), survival (time to CLABSI) and competing risks (time to CLABSI, discharge or death) outcomes to predict the 7-day CLABSI risk. We evaluated model performance across 100 train/test splits. Performance of binary, multinomial and competing risks models was similar: AUROC was 0.74 for baseline predictions, rose to 0.78 for predictions at day 5 in the catheter episode, and decreased thereafter. Survival models overestimated the risk of CLABSI (E:O ratios between 1.2 and 1.6), and had AUROCs about 0.01 lower than other models. Binary and multinomial models had lowest computation times. Models including multiple outcome events (multinomial and competing risks) display a different internal structure compared to binary and survival models. In the absence of censoring, complex modelling choices do not considerably improve the predictive performance compared to a binary model for CLABSI prediction in our studied settings. Survival models censoring the competing events at their time of occurrence should be avoided.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge