Keith Steele
DASGAN -- Joint Domain Adaptation and Segmentation for the Analysis of Epithelial Regions in Histopathology PD-L1 Images
Jun 26, 2019



Abstract:The analysis of the tumor environment on digital histopathology slides is becoming key for the understanding of the immune response against cancer, supporting the development of novel immuno-therapies. We introduce here a novel deep learning solution to the related problem of tumor epithelium segmentation. While most existing deep learning segmentation approaches are trained on time-consuming and costly manual annotation on single stain domain (PD-L1), we leverage here semi-automatically labeled images from a second stain domain (Cytokeratin-CK). We introduce an end-to-end trainable network that jointly segment tumor epithelium on PD-L1 while leveraging unpaired image-to-image translation between CK and PD-L1, therefore completely bypassing the need for serial sections or re-staining of slides. Extending the method to differentiate between PD-L1 positive and negative tumor epithelium regions enables the automated estimation of the PD-L1 Tumor Cell (TC) score. Quantitative experimental results demonstrate the accuracy of our approach against state-of-the-art segmentation methods.
Deep Semi Supervised Generative Learning for Automated PD-L1 Tumor Cell Scoring on NSCLC Tissue Needle Biopsies
Jun 28, 2018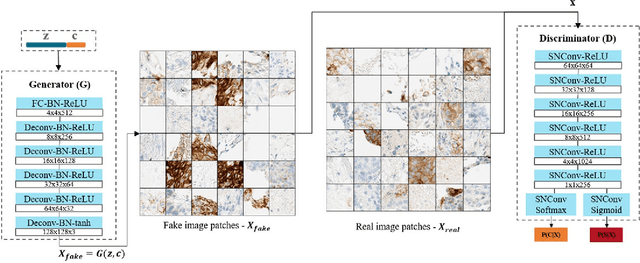
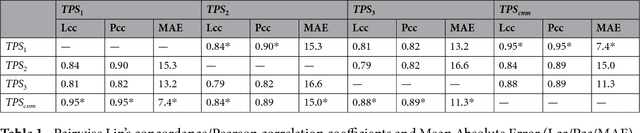

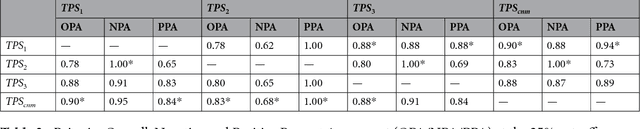
Abstract:The level of PD-L1 expression in immunohistochemistry (IHC) assays is a key biomarker for the identification of Non-Small-Cell-Lung-Cancer (NSCLC) patients that may respond to anti PD-1/PD-L1 treatments. The quantification of PD-L1 expression currently includes the visual estimation of a Tumor Cell (TC) score by a pathologist and consists of evaluating the ratio of PD-L1 positive and PD-L1 negative tumor cells. Known challenges like differences in positivity estimation around clinically relevant cut-offs and sub-optimal quality of samples makes visual scoring tedious and subjective, yielding a scoring variability between pathologists. In this work, we propose a novel deep learning solution that enables the first automated and objective scoring of PD-L1 expression in late stage NSCLC needle biopsies. To account for the low amount of tissue available in biopsy images and to restrict the amount of manual annotations necessary for training, we explore the use of semi-supervised approaches against standard fully supervised methods. We consolidate the manual annotations used for training as well the visual TC scores used for quantitative evaluation with multiple pathologists. Concordance measures computed on a set of slides unseen during training provide evidence that our automatic scoring method matches visual scoring on the considered dataset while ensuring repeatability and objectivity.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge