Kara M. Kelly
Automatic Quantification of Serial PET/CT Images for Pediatric Hodgkin Lymphoma Patients Using a Longitudinally-Aware Segmentation Network
Apr 12, 2024


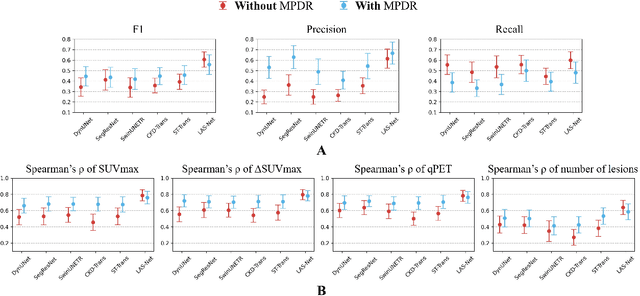
Abstract:$\textbf{Purpose}$: Automatic quantification of longitudinal changes in PET scans for lymphoma patients has proven challenging, as residual disease in interim-therapy scans is often subtle and difficult to detect. Our goal was to develop a longitudinally-aware segmentation network (LAS-Net) that can quantify serial PET/CT images for pediatric Hodgkin lymphoma patients. $\textbf{Materials and Methods}$: This retrospective study included baseline (PET1) and interim (PET2) PET/CT images from 297 patients enrolled in two Children's Oncology Group clinical trials (AHOD1331 and AHOD0831). LAS-Net incorporates longitudinal cross-attention, allowing relevant features from PET1 to inform the analysis of PET2. Model performance was evaluated using Dice coefficients for PET1 and detection F1 scores for PET2. Additionally, we extracted and compared quantitative PET metrics, including metabolic tumor volume (MTV) and total lesion glycolysis (TLG) in PET1, as well as qPET and $\Delta$SUVmax in PET2, against physician measurements. We quantified their agreement using Spearman's $\rho$ correlations and employed bootstrap resampling for statistical analysis. $\textbf{Results}$: LAS-Net detected residual lymphoma in PET2 with an F1 score of 0.606 (precision/recall: 0.615/0.600), outperforming all comparator methods (P<0.01). For baseline segmentation, LAS-Net achieved a mean Dice score of 0.772. In PET quantification, LAS-Net's measurements of qPET, $\Delta$SUVmax, MTV and TLG were strongly correlated with physician measurements, with Spearman's $\rho$ of 0.78, 0.80, 0.93 and 0.96, respectively. The performance remained high, with a slight decrease, in an external testing cohort. $\textbf{Conclusion}$: LAS-Net achieved high performance in quantifying PET metrics across serial scans, highlighting the value of longitudinal awareness in evaluating multi-time-point imaging datasets.
Automatic Personalized Impression Generation for PET Reports Using Large Language Models
Sep 18, 2023Abstract:Purpose: To determine if fine-tuned large language models (LLMs) can generate accurate, personalized impressions for whole-body PET reports. Materials and Methods: Twelve language models were trained on a corpus of PET reports using the teacher-forcing algorithm, with the report findings as input and the clinical impressions as reference. An extra input token encodes the reading physician's identity, allowing models to learn physician-specific reporting styles. Our corpus comprised 37,370 retrospective PET reports collected from our institution between 2010 and 2022. To identify the best LLM, 30 evaluation metrics were benchmarked against quality scores from two nuclear medicine (NM) physicians, with the most aligned metrics selecting the model for expert evaluation. In a subset of data, model-generated impressions and original clinical impressions were assessed by three NM physicians according to 6 quality dimensions and an overall utility score (5-point scale). Each physician reviewed 12 of their own reports and 12 reports from other physicians. Bootstrap resampling was used for statistical analysis. Results: Of all evaluation metrics, domain-adapted BARTScore and PEGASUSScore showed the highest Spearman's rho correlations (0.568 and 0.563) with physician preferences. Based on these metrics, the fine-tuned PEGASUS model was selected as the top LLM. When physicians reviewed PEGASUS-generated impressions in their own style, 89% were considered clinically acceptable, with a mean utility score of 4.08/5. Physicians rated these personalized impressions as comparable in overall utility to the impressions dictated by other physicians (4.03, P=0.41). Conclusion: Personalized impressions generated by PEGASUS were clinically useful, highlighting its potential to expedite PET reporting.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge