Joshua A. Rackers
Strain Problems got you in a Twist? Try StrainRelief: A Quantum-Accurate Tool for Ligand Strain Calculations
Mar 17, 2025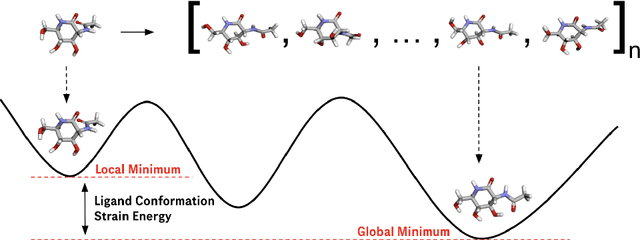
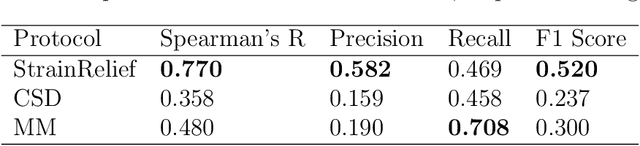


Abstract:Ligand strain energy, the energy difference between the bound and unbound conformations of a ligand, is an important component of structure-based small molecule drug design. A large majority of observed ligands in protein-small molecule co-crystal structures bind in low-strain conformations, making strain energy a useful filter for structure-based drug design. In this work we present a tool for calculating ligand strain with a high accuracy. StrainRelief uses a MACE Neural Network Potential (NNP), trained on a large database of Density Functional Theory (DFT) calculations to estimate ligand strain of neutral molecules with quantum accuracy. We show that this tool estimates strain energy differences relative to DFT to within 1.4 kcal/mol, more accurately than alternative NNPs. These results highlight the utility of NNPs in drug discovery, and provide a useful tool for drug discovery teams.
On the design space between molecular mechanics and machine learning force fields
Sep 03, 2024Abstract:A force field as accurate as quantum mechanics (QM) and as fast as molecular mechanics (MM), with which one can simulate a biomolecular system efficiently enough and meaningfully enough to get quantitative insights, is among the most ardent dreams of biophysicists -- a dream, nevertheless, not to be fulfilled any time soon. Machine learning force fields (MLFFs) represent a meaningful endeavor towards this direction, where differentiable neural functions are parametrized to fit ab initio energies, and furthermore forces through automatic differentiation. We argue that, as of now, the utility of the MLFF models is no longer bottlenecked by accuracy but primarily by their speed (as well as stability and generalizability), as many recent variants, on limited chemical spaces, have long surpassed the chemical accuracy of $1$ kcal/mol -- the empirical threshold beyond which realistic chemical predictions are possible -- though still magnitudes slower than MM. Hoping to kindle explorations and designs of faster, albeit perhaps slightly less accurate MLFFs, in this review, we focus our attention on the design space (the speed-accuracy tradeoff) between MM and ML force fields. After a brief review of the building blocks of force fields of either kind, we discuss the desired properties and challenges now faced by the force field development community, survey the efforts to make MM force fields more accurate and ML force fields faster, envision what the next generation of MLFF might look like.
Hierarchical Learning in Euclidean Neural Networks
Oct 10, 2022



Abstract:Equivariant machine learning methods have shown wide success at 3D learning applications in recent years. These models explicitly build in the reflection, translation and rotation symmetries of Euclidean space and have facilitated large advances in accuracy and data efficiency for a range of applications in the physical sciences. An outstanding question for equivariant models is why they achieve such larger-than-expected advances in these applications. To probe this question, we examine the role of higher order (non-scalar) features in Euclidean Neural Networks (\texttt{e3nn}). We focus on the previously studied application of \texttt{e3nn} to the problem of electron density prediction, which allows for a variety of non-scalar outputs, and examine whether the nature of the output (scalar $l=0$, vector $l=1$, or higher order $l>1$) is relevant to the effectiveness of non-scalar hidden features in the network. Further, we examine the behavior of non-scalar features throughout training, finding a natural hierarchy of features by $l$, reminiscent of a multipole expansion. We aim for our work to ultimately inform design principles and choices of domain applications for {\tt e3nn} networks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge