Johann A. Briffa
Multitemporal and multispectral data fusion for super-resolution of Sentinel-2 images
Jan 26, 2023

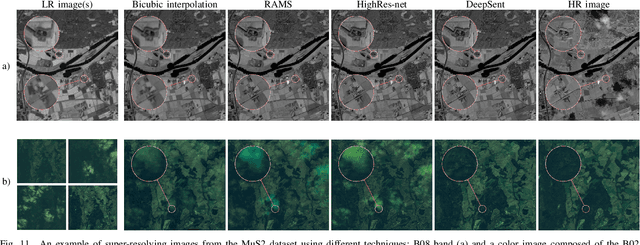

Abstract:Multispectral Sentinel-2 images are a valuable source of Earth observation data, however spatial resolution of their spectral bands limited to 10 m, 20 m, and 60 m ground sampling distance remains insufficient in many cases. This problem can be addressed with super-resolution, aimed at reconstructing a high-resolution image from a low-resolution observation. For Sentinel-2, spectral information fusion allows for enhancing the 20 m and 60 m bands to the 10 m resolution. Also, there were attempts to combine multitemporal stacks of individual Sentinel-2 bands, however these two approaches have not been combined so far. In this paper, we introduce DeepSent -- a new deep network for super-resolving multitemporal series of multispectral Sentinel-2 images. It is underpinned with information fusion performed simultaneously in the spectral and temporal dimensions to generate an enlarged multispectral image. In our extensive experimental study, we demonstrate that our solution outperforms other state-of-the-art techniques that realize either multitemporal or multispectral data fusion. Furthermore, we show that the advantage of DeepSent results from how these two fusion types are combined in a single architecture, which is superior to performing such fusion in a sequential manner. Importantly, we have applied our method to super-resolve real-world Sentinel-2 images, enhancing the spatial resolution of all the spectral bands to 3.3 m nominal ground sampling distance, and we compare the outcome with very high-resolution WorldView-2 images. We will publish our implementation upon paper acceptance, and we expect it will increase the possibilities of exploiting super-resolved Sentinel-2 images in real-life applications.
Automated segmentation of microtomography imaging of Egyptian mummies
May 14, 2021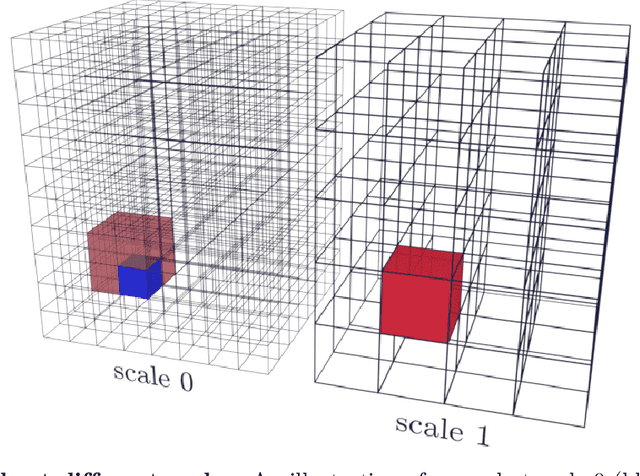
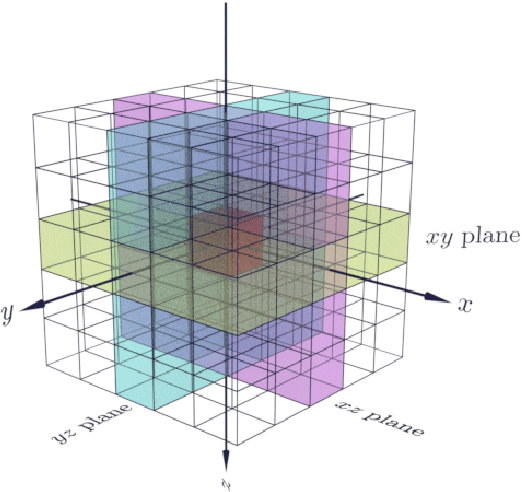
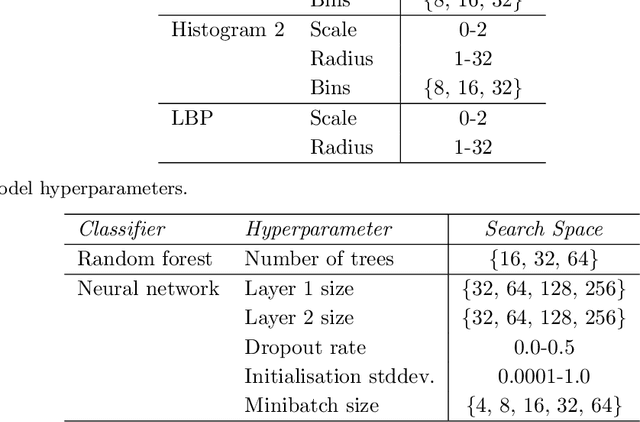

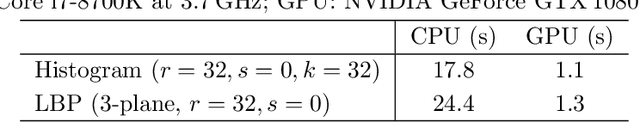
Abstract:Propagation Phase Contrast Synchrotron Microtomography (PPC-SR${\mu}$CT) is the gold standard for non-invasive and non-destructive access to internal structures of archaeological remains. In this analysis, the virtual specimen needs to be segmented to separate different parts or materials, a process that normally requires considerable human effort. In the Automated SEgmentation of Microtomography Imaging (ASEMI) project, we developed a tool to automatically segment these volumetric images, using manually segmented samples to tune and train a machine learning model. For a set of four specimens of ancient Egyptian animal mummies we achieve an overall accuracy of 94-98% when compared with manually segmented slices, approaching the results of off-the-shelf commercial software using deep learning (97-99%) at much lower complexity. A qualitative analysis of the segmented output shows that our results are close in term of usability to those from deep learning, justifying the use of these techniques.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge