Jiyoon Yi
Enhancing AI microscopy for foodborne bacterial classification via adversarial domain adaptation across optical and biological variability
Nov 29, 2024Abstract:Rapid detection of foodborne bacteria is critical for food safety and quality, yet traditional culture-based methods require extended incubation and specialized sample preparation. This study addresses these challenges by i) enhancing the generalizability of AI-enabled microscopy for bacterial classification using adversarial domain adaptation and ii) comparing the performance of single-target and multi-domain adaptation. Three Gram-positive (Bacillus coagulans, Bacillus subtilis, Listeria innocua) and three Gram-negative (E. coli, Salmonella Enteritidis, Salmonella Typhimurium) strains were classified. EfficientNetV2 served as the backbone architecture, leveraging fine-grained feature extraction for small targets. Few-shot learning enabled scalability, with domain-adversarial neural networks (DANNs) addressing single domains and multi-DANNs (MDANNs) generalizing across all target domains. The model was trained on source domain data collected under controlled conditions (phase contrast microscopy, 60x magnification, 3-h bacterial incubation) and evaluated on target domains with variations in microscopy modality (brightfield, BF), magnification (20x), and extended incubation to compensate for lower resolution (20x-5h). DANNs improved target domain classification accuracy by up to 54.45% (20x), 43.44% (20x-5h), and 31.67% (BF), with minimal source domain degradation (<4.44%). MDANNs achieved superior performance in the BF domain and substantial gains in the 20x domain. Grad-CAM and t-SNE visualizations validated the model's ability to learn domain-invariant features across diverse conditions. This study presents a scalable and adaptable framework for bacterial classification, reducing reliance on extensive sample preparation and enabling application in decentralized and resource-limited environments.
Label-free prediction of fluorescence markers in bovine satellite cells using deep learning
Oct 17, 2024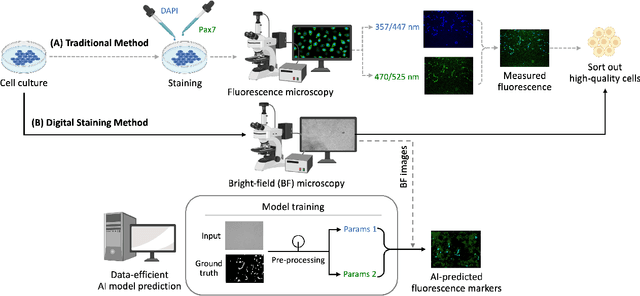

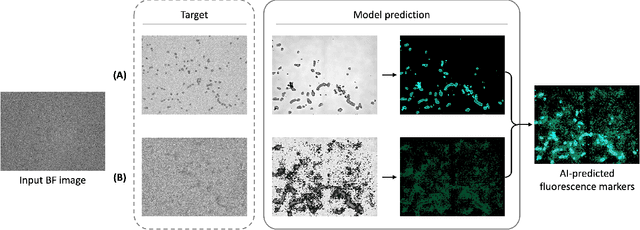
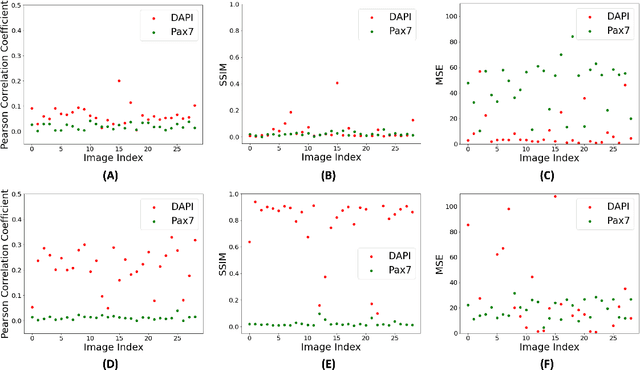
Abstract:Assessing the quality of bovine satellite cells (BSCs) is essential for the cultivated meat industry, which aims to address global food sustainability challenges. This study aims to develop a label-free method for predicting fluorescence markers in isolated BSCs using deep learning. We employed a U-Net-based CNN model to predict multiple fluorescence signals from a single bright-field microscopy image of cell culture. Two key biomarkers, DAPI and Pax7, were used to determine the abundance and quality of BSCs. The image pre-processing pipeline included fluorescence denoising to improve prediction performance and consistency. A total of 48 biological replicates were used, with statistical performance metrics such as Pearson correlation coefficient and SSIM employed for model evaluation. The model exhibited better performance with DAPI predictions due to uniform staining. Pax7 predictions were more variable, reflecting biological heterogeneity. Enhanced visualization techniques, including color mapping and image overlay, improved the interpretability of the predictions by providing better contextual and perceptual information. The findings highlight the importance of data pre-processing and demonstrate the potential of deep learning to advance non-invasive, label-free assessment techniques in the cultivated meat industry, paving the way for reliable and actionable AI-driven evaluations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge