Ilse Slootweg
Report-Guided Automatic Lesion Annotation for Deep Learning-Based Prostate Cancer Detection in bpMRI
Dec 09, 2021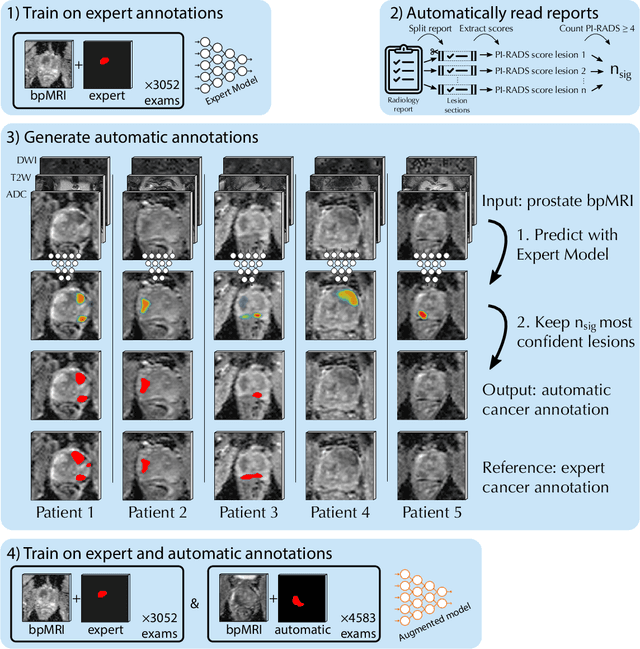

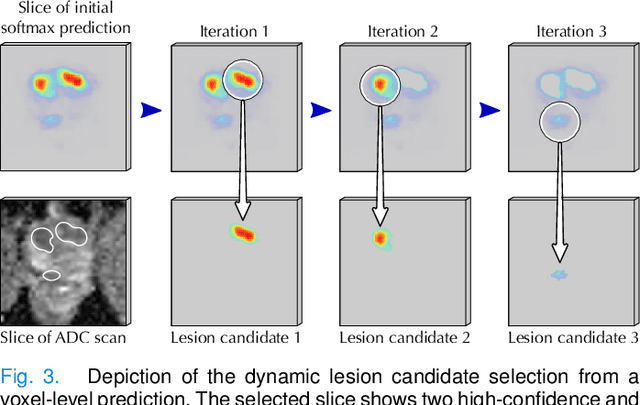
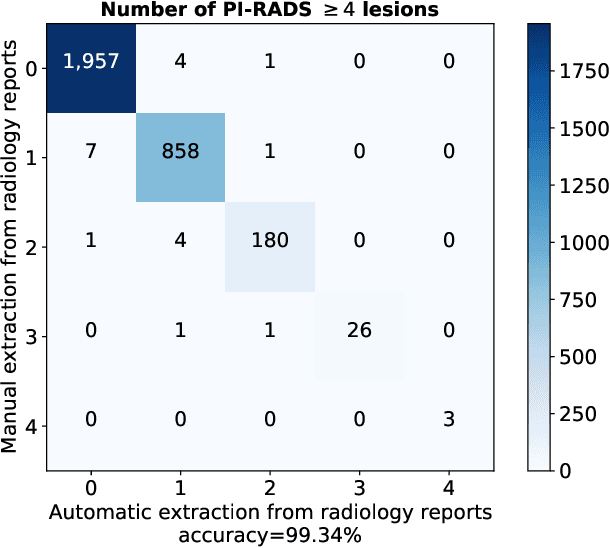
Abstract:Deep learning-based diagnostic performance increases with more annotated data, but manual annotation is a bottleneck in most fields. Experts evaluate diagnostic images during clinical routine, and write their findings in reports. Automatic annotation based on clinical reports could overcome the manual labelling bottleneck. We hypothesise that dense annotations for detection tasks can be generated using model predictions, guided by sparse information from these reports. To demonstrate efficacy, we generated clinically significant prostate cancer (csPCa) annotations, guided by the number of clinically significant findings in the radiology reports. We included 7,756 prostate MRI examinations, of which 3,050 were manually annotated and 4,706 were automatically annotated. We evaluated the automatic annotation quality on the manually annotated subset: our score extraction correctly identified the number of csPCa lesions for $99.3\%$ of the reports and our csPCa segmentation model correctly localised $83.8 \pm 1.1\%$ of the lesions. We evaluated prostate cancer detection performance on 300 exams from an external centre with histopathology-confirmed ground truth. Augmenting the training set with automatically labelled exams improved patient-based diagnostic area under the receiver operating characteristic curve from $88.1\pm 1.1\%$ to $89.8\pm 1.0\%$ ($P = 1.2 \cdot 10^{-4}$) and improved lesion-based sensitivity at one false positive per case from $79.2 \pm 2.8\%$ to $85.4 \pm 1.9\%$ ($P<10^{-4}$), with $mean \pm std.$ over 15 independent runs. This improved performance demonstrates the feasibility of our report-guided automatic annotations. Source code is made publicly available at https://github.com/DIAGNijmegen/Report-Guided-Annotation. Best csPCa detection algorithm is made available at https://grand-challenge.org/algorithms/bpmri-cspca-detection-report-guided-annotations/.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge