Fabian Braeu
Are Macula or Optic Nerve Head Structures better at Diagnosing Glaucoma? An Answer using AI and Wide-Field Optical Coherence Tomography
Oct 13, 2022
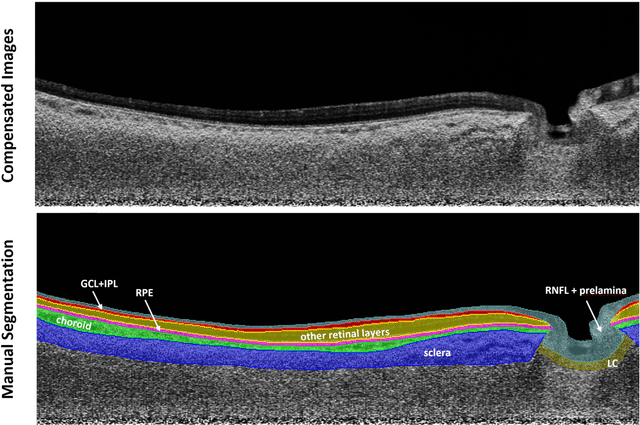
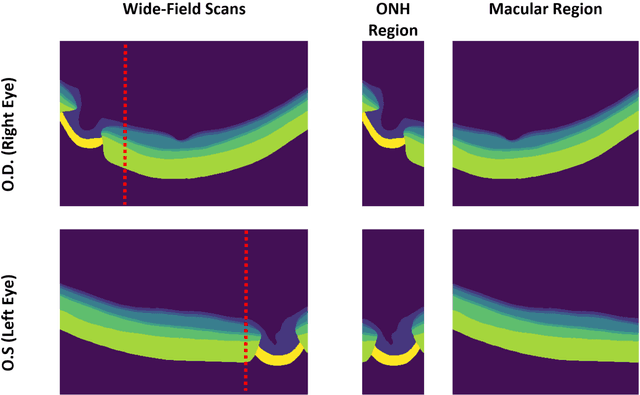
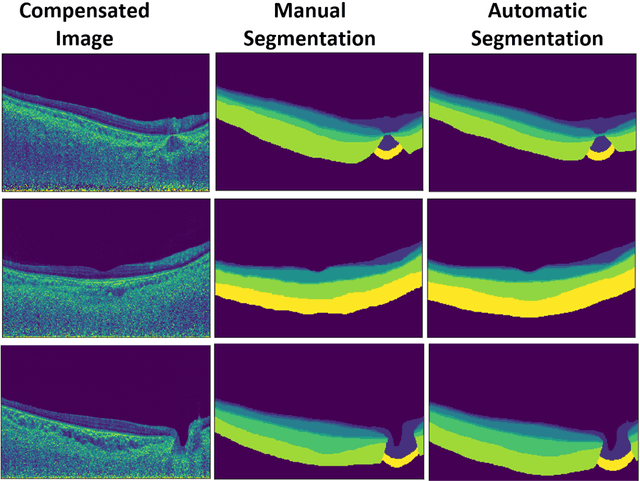
Abstract:Purpose: (1) To develop a deep learning algorithm to automatically segment structures of the optic nerve head (ONH) and macula in 3D wide-field optical coherence tomography (OCT) scans; (2) To assess whether 3D macula or ONH structures (or the combination of both) provide the best diagnostic power for glaucoma. Methods: A cross-sectional comparative study was performed which included wide-field swept-source OCT scans from 319 glaucoma subjects and 298 non-glaucoma subjects. All scans were compensated to improve deep-tissue visibility. We developed a deep learning algorithm to automatically label all major ONH tissue structures by using 270 manually annotated B-scans for training. The performance of our algorithm was assessed using the Dice coefficient (DC). A glaucoma classification algorithm (3D CNN) was then designed using a combination of 500 OCT volumes and their corresponding automatically segmented masks. This algorithm was trained and tested on 3 datasets: OCT scans cropped to contain the macular tissues only, those to contain the ONH tissues only, and the full wide-field OCT scans. The classification performance for each dataset was reported using the AUC. Results: Our segmentation algorithm was able to segment ONH and macular tissues with a DC of 0.94 $\pm$ 0.003. The classification algorithm was best able to diagnose glaucoma using wide-field 3D-OCT volumes with an AUC of 0.99 $\pm$ 0.01, followed by ONH volumes with an AUC of 0.93 $\pm$ 0.06, and finally macular volumes with an AUC of 0.91 $\pm$ 0.11. Conclusions: this study showed that using wide-field OCT as compared to the typical OCT images containing just the ONH or macular may allow for a significantly improved glaucoma diagnosis. This may encourage the mainstream adoption of 3D wide-field OCT scans. For clinical AI studies that use traditional machines, we would recommend the use of ONH scans as opposed to macula scans.
Medical Application of Geometric Deep Learning for the Diagnosis of Glaucoma
Apr 14, 2022
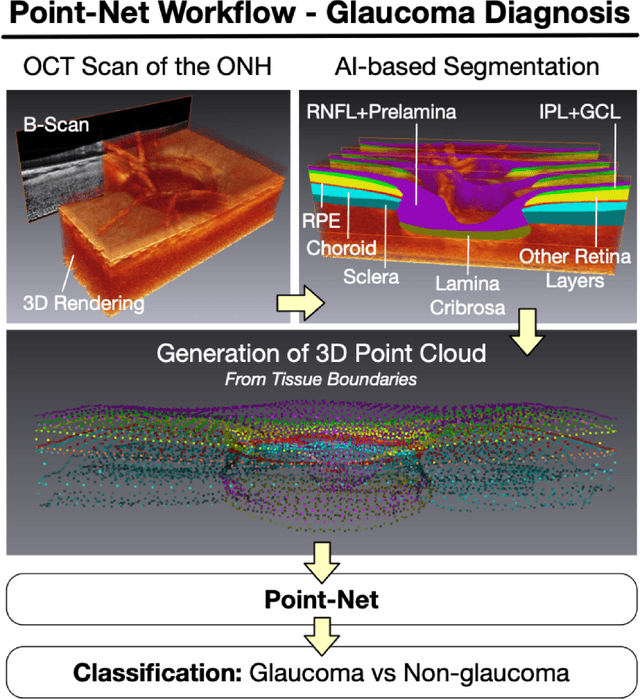
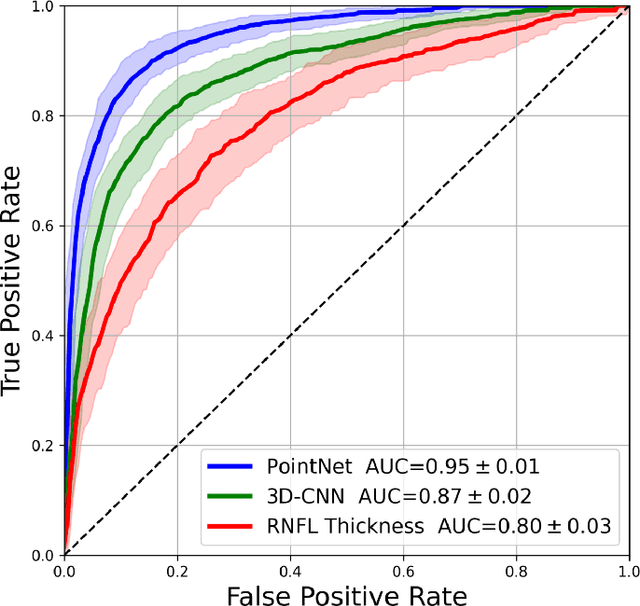
Abstract:Purpose: (1) To assess the performance of geometric deep learning (PointNet) in diagnosing glaucoma from a single optical coherence tomography (OCT) 3D scan of the optic nerve head (ONH); (2) To compare its performance to that obtained with a standard 3D convolutional neural network (CNN), and with a gold-standard glaucoma parameter, i.e. retinal nerve fiber layer (RNFL) thickness. Methods: 3D raster scans of the ONH were acquired with Spectralis OCT for 477 glaucoma and 2,296 non-glaucoma subjects at the Singapore National Eye Centre. All volumes were automatically segmented using deep learning to identify 7 major neural and connective tissues including the RNFL, the prelamina, and the lamina cribrosa (LC). Each ONH was then represented as a 3D point cloud with 1,000 points chosen randomly from all tissue boundaries. To simplify the problem, all ONH point clouds were aligned with respect to the plane and center of Bruch's membrane opening. Geometric deep learning (PointNet) was then used to provide a glaucoma diagnosis from a single OCT point cloud. The performance of our approach was compared to that obtained with a 3D CNN, and with RNFL thickness. Results: PointNet was able to provide a robust glaucoma diagnosis solely from the ONH represented as a 3D point cloud (AUC=95%). The performance of PointNet was superior to that obtained with a standard 3D CNN (AUC=87%) and with that obtained from RNFL thickness alone (AUC=80%). Discussion: We provide a proof-of-principle for the application of geometric deep learning in the field of glaucoma. Our technique requires significantly less information as input to perform better than a 3D CNN, and with an AUC superior to that obtained from RNFL thickness alone. Geometric deep learning may have wide applicability in the field of Ophthalmology.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge