Diana Montezuma
CountPath: Automating Fragment Counting in Digital Pathology
Mar 13, 2025Abstract:Quality control of medical images is a critical component of digital pathology, ensuring that diagnostic images meet required standards. A pre-analytical task within this process is the verification of the number of specimen fragments, a process that ensures that the number of fragments on a slide matches the number documented in the macroscopic report. This step is important to ensure that the slides contain the appropriate diagnostic material from the grossing process, thereby guaranteeing the accuracy of subsequent microscopic examination and diagnosis. Traditionally, this assessment is performed manually, requiring significant time and effort while being subject to significant variability due to its subjective nature. To address these challenges, this study explores an automated approach to fragment counting using the YOLOv9 and Vision Transformer models. Our results demonstrate that the automated system achieves a level of performance comparable to expert assessments, offering a reliable and efficient alternative to manual counting. Additionally, we present findings on interobserver variability, showing that the automated approach achieves an accuracy of 86%, which falls within the range of variation observed among experts (82-88%), further supporting its potential for integration into routine pathology workflows.
A Survey on Cell Nuclei Instance Segmentation and Classification: Leveraging Context and Attention
Jul 26, 2024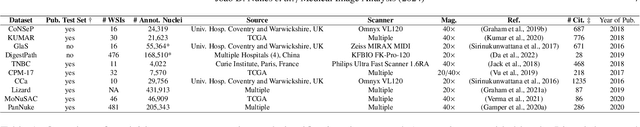
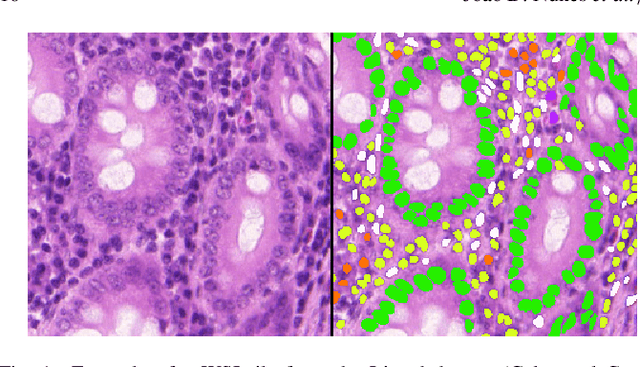
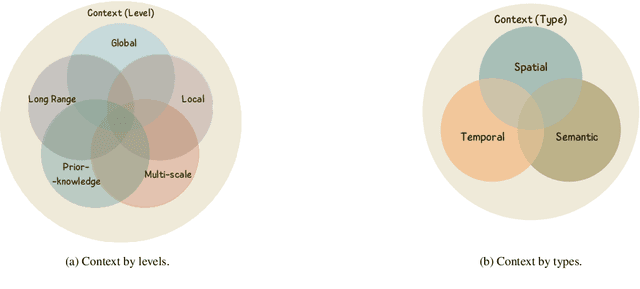
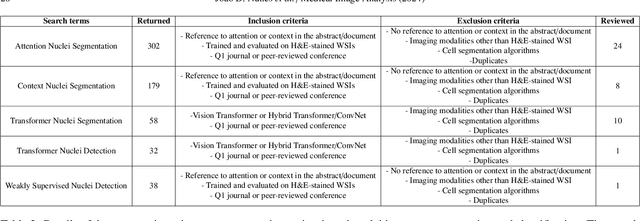
Abstract:Manually annotating nuclei from the gigapixel Hematoxylin and Eosin (H&E)-stained Whole Slide Images (WSIs) is a laborious and costly task, meaning automated algorithms for cell nuclei instance segmentation and classification could alleviate the workload of pathologists and clinical researchers and at the same time facilitate the automatic extraction of clinically interpretable features. But due to high intra- and inter-class variability of nuclei morphological and chromatic features, as well as H&E-stains susceptibility to artefacts, state-of-the-art algorithms cannot correctly detect and classify instances with the necessary performance. In this work, we hypothesise context and attention inductive biases in artificial neural networks (ANNs) could increase the generalization of algorithms for cell nuclei instance segmentation and classification. We conduct a thorough survey on context and attention methods for cell nuclei instance segmentation and classification from H&E-stained microscopy imaging, while providing a comprehensive discussion of the challenges being tackled with context and attention. Besides, we illustrate some limitations of current approaches and present ideas for future research. As a case study, we extend both a general instance segmentation and classification method (Mask-RCNN) and a tailored cell nuclei instance segmentation and classification model (HoVer-Net) with context- and attention-based mechanisms, and do a comparative analysis on a multi-centre colon nuclei identification and counting dataset. Although pathologists rely on context at multiple levels while paying attention to specific Regions of Interest (RoIs) when analysing and annotating WSIs, our findings suggest translating that domain knowledge into algorithm design is no trivial task, but to fully exploit these mechanisms, the scientific understanding of these methods should be addressed.
A CAD System for Colorectal Cancer from WSI: A Clinically Validated Interpretable ML-based Prototype
Jan 06, 2023Abstract:The integration of Artificial Intelligence (AI) and Digital Pathology has been increasing over the past years. Nowadays, applications of deep learning (DL) methods to diagnose cancer from whole-slide images (WSI) are, more than ever, a reality within different research groups. Nonetheless, the development of these systems was limited by a myriad of constraints regarding the lack of training samples, the scaling difficulties, the opaqueness of DL methods, and, more importantly, the lack of clinical validation. As such, we propose a system designed specifically for the diagnosis of colorectal samples. The construction of such a system consisted of four stages: (1) a careful data collection and annotation process, which resulted in one of the largest WSI colorectal samples datasets; (2) the design of an interpretable mixed-supervision scheme to leverage the domain knowledge introduced by pathologists through spatial annotations; (3) the development of an effective sampling approach based on the expected severeness of each tile, which decreased the computation cost by a factor of almost 6x; (4) the creation of a prototype that integrates the full set of features of the model to be evaluated in clinical practice. During these stages, the proposed method was evaluated in four separate test sets, two of them are external and completely independent. On the largest of those sets, the proposed approach achieved an accuracy of 93.44%. DL for colorectal samples is a few steps closer to stop being research exclusive and to become fully integrated in clinical practice.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge