David S. Lim
Bridging the gap between prostate radiology and pathology through machine learning
Dec 03, 2021
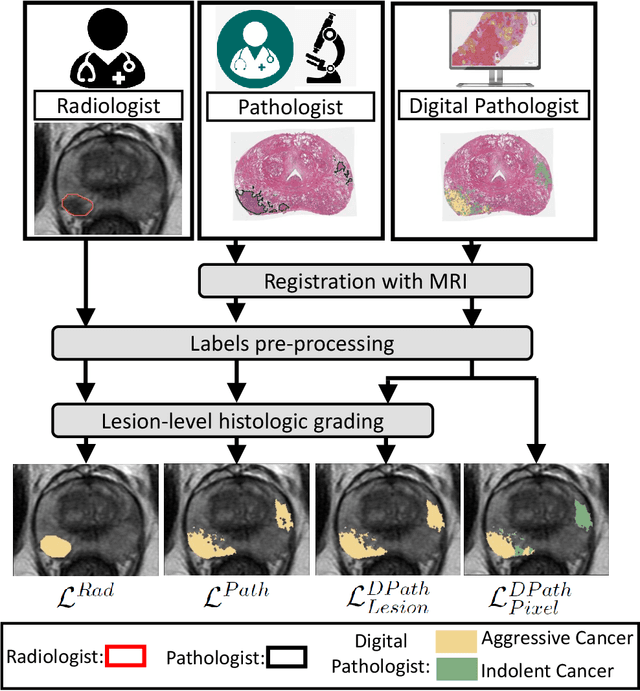
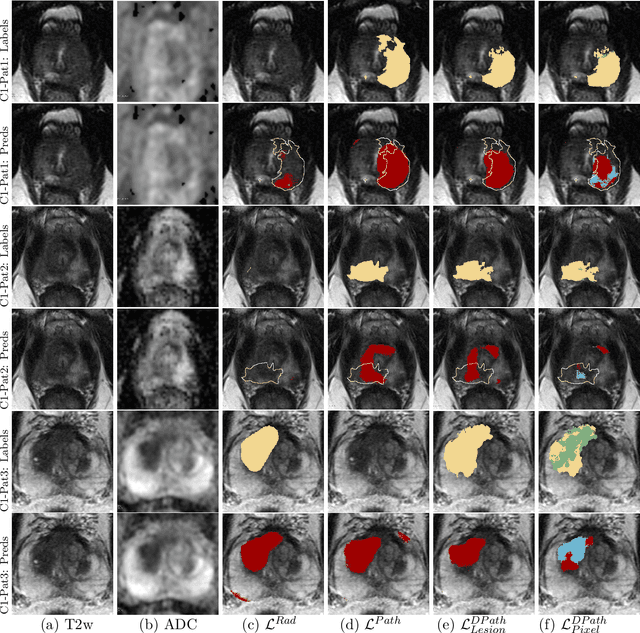
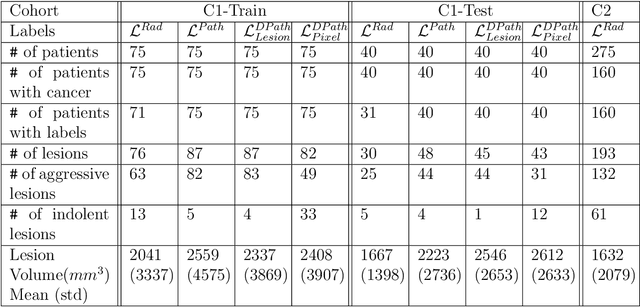
Abstract:Prostate cancer is the second deadliest cancer for American men. While Magnetic Resonance Imaging (MRI) is increasingly used to guide targeted biopsies for prostate cancer diagnosis, its utility remains limited due to high rates of false positives and false negatives as well as low inter-reader agreements. Machine learning methods to detect and localize cancer on prostate MRI can help standardize radiologist interpretations. However, existing machine learning methods vary not only in model architecture, but also in the ground truth labeling strategies used for model training. In this study, we compare different labeling strategies, namely, pathology-confirmed radiologist labels, pathologist labels on whole-mount histopathology images, and lesion-level and pixel-level digital pathologist labels (previously validated deep learning algorithm on histopathology images to predict pixel-level Gleason patterns) on whole-mount histopathology images. We analyse the effects these labels have on the performance of the trained machine learning models. Our experiments show that (1) radiologist labels and models trained with them can miss cancers, or underestimate cancer extent, (2) digital pathologist labels and models trained with them have high concordance with pathologist labels, and (3) models trained with digital pathologist labels achieve the best performance in prostate cancer detection in two different cohorts with different disease distributions, irrespective of the model architecture used. Digital pathologist labels can reduce challenges associated with human annotations, including labor, time, inter- and intra-reader variability, and can help bridge the gap between prostate radiology and pathology by enabling the training of reliable machine learning models to detect and localize prostate cancer on MRI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge