Craig A. Glastonbury
Adjusting for Confounding in Unsupervised Latent Representations of Images
Nov 26, 2018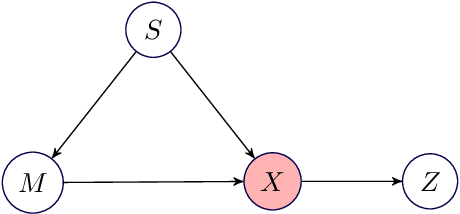
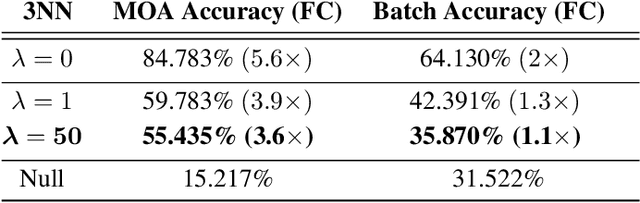
Abstract:Biological imaging data are often partially confounded or contain unwanted variability. Examples of such phenomena include variable lighting across microscopy image captures, stain intensity variation in histological slides, and batch effects for high throughput drug screening assays. Therefore, to develop "fair" models which generalise well to unseen examples, it is crucial to learn data representations that are insensitive to nuisance factors of variation. In this paper, we present a strategy based on adversarial training, capable of learning unsupervised representations invariant to confounders. As an empirical validation of our method, we use deep convolutional autoencoders to learn unbiased cellular representations from microscopy imaging.
Towards Deep Cellular Phenotyping in Placental Histology
May 25, 2018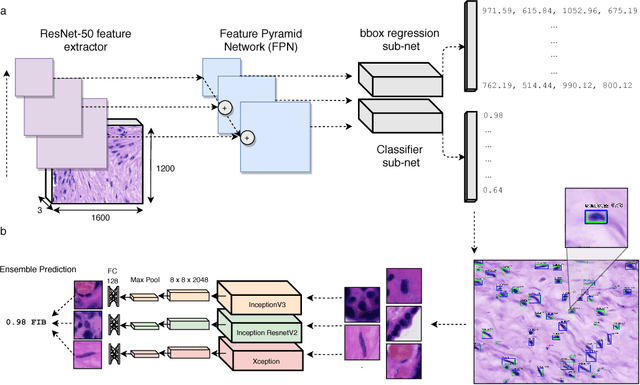
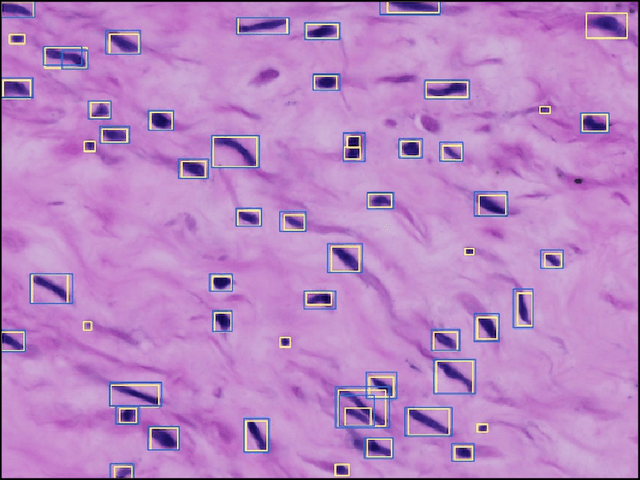
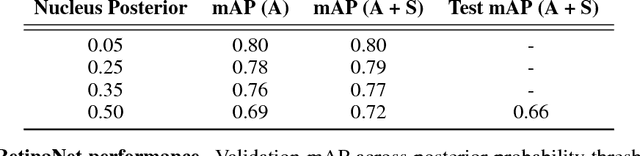

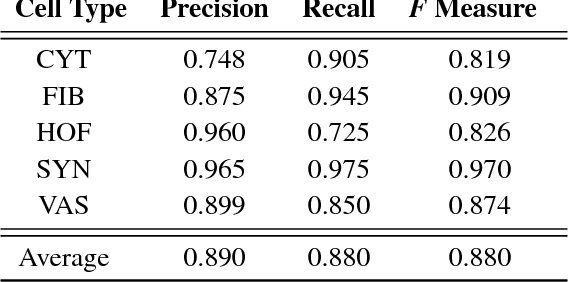
Abstract:The placenta is a complex organ, playing multiple roles during fetal development. Very little is known about the association between placental morphological abnormalities and fetal physiology. In this work, we present an open sourced, computationally tractable deep learning pipeline to analyse placenta histology at the level of the cell. By utilising two deep Convolutional Neural Network architectures and transfer learning, we can robustly localise and classify placental cells within five classes with an accuracy of 89%. Furthermore, we learn deep embeddings encoding phenotypic knowledge that is capable of both stratifying five distinct cell populations and learn intraclass phenotypic variance. We envisage that the automation of this pipeline to population scale studies of placenta histology has the potential to improve our understanding of basic cellular placental biology and its variations, particularly its role in predicting adverse birth outcomes.
Count-ception: Counting by Fully Convolutional Redundant Counting
Jul 23, 2017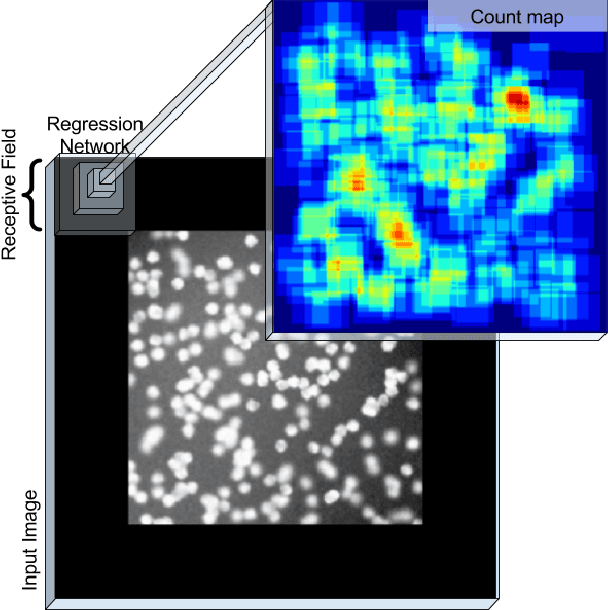
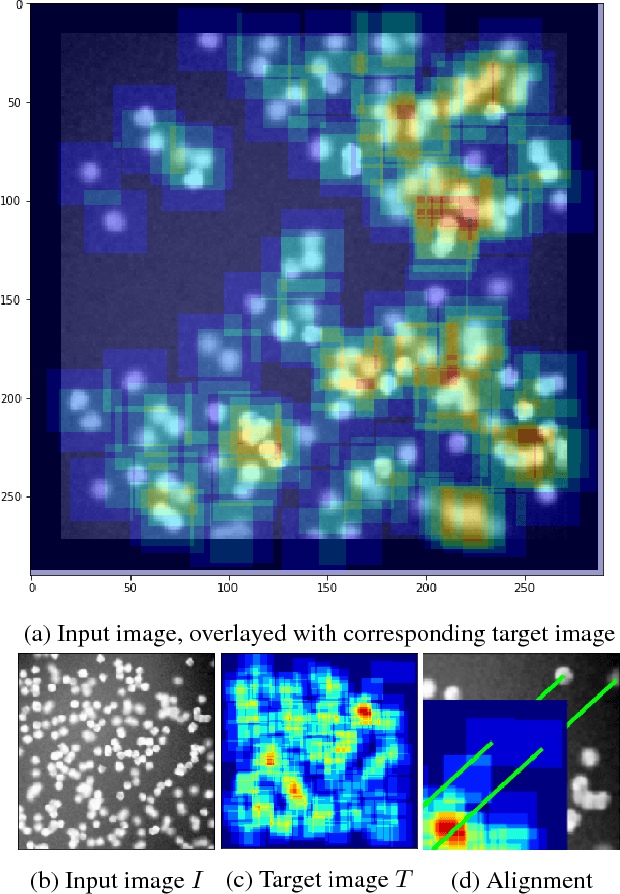
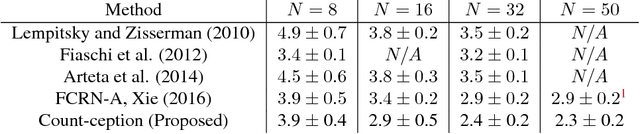
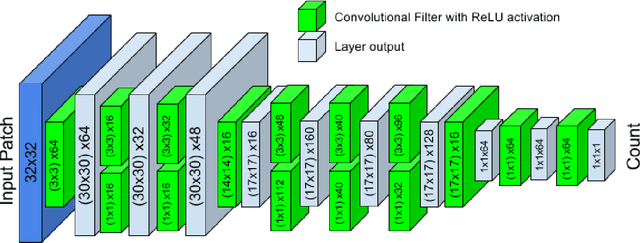
Abstract:Counting objects in digital images is a process that should be replaced by machines. This tedious task is time consuming and prone to errors due to fatigue of human annotators. The goal is to have a system that takes as input an image and returns a count of the objects inside and justification for the prediction in the form of object localization. We repose a problem, originally posed by Lempitsky and Zisserman, to instead predict a count map which contains redundant counts based on the receptive field of a smaller regression network. The regression network predicts a count of the objects that exist inside this frame. By processing the image in a fully convolutional way each pixel is going to be accounted for some number of times, the number of windows which include it, which is the size of each window, (i.e., 32x32 = 1024). To recover the true count we take the average over the redundant predictions. Our contribution is redundant counting instead of predicting a density map in order to average over errors. We also propose a novel deep neural network architecture adapted from the Inception family of networks called the Count-ception network. Together our approach results in a 20% relative improvement (2.9 to 2.3 MAE) over the state of the art method by Xie, Noble, and Zisserman in 2016.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge