Chaitra V. Hegde
Knee Cartilage Segmentation Using Diffusion-Weighted MRI
Dec 04, 2019
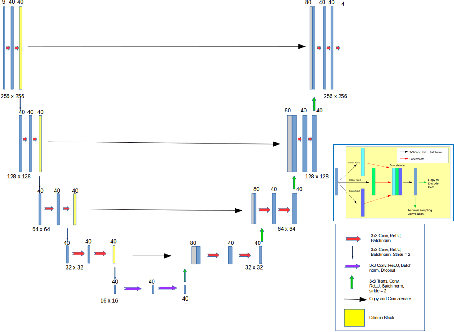
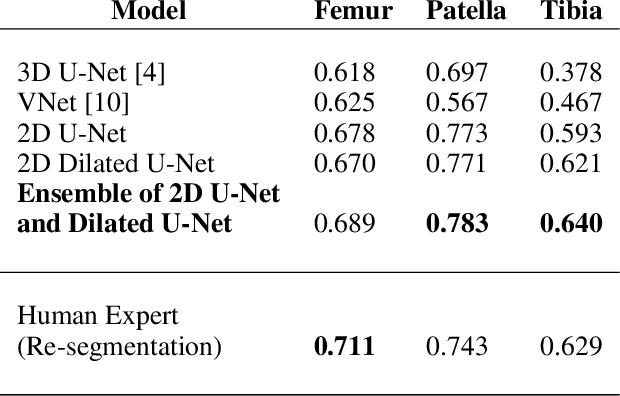
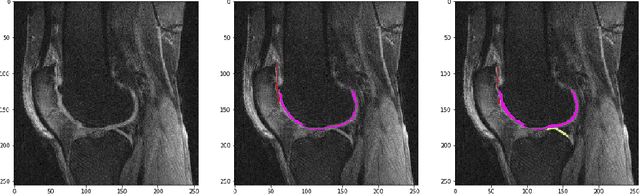
Abstract:The integrity of articular cartilage is a crucial aspect in the early diagnosis of osteoarthritis (OA). Many novel MRI techniques have the potential to assess compositional changes of the cartilage extracellular matrix. Among these techniques, diffusion tensor imaging (DTI) of cartilage provides a simultaneous assessment of the two principal components of the solid matrix: collagen structure and proteoglycan concentration. DTI, as for any other compositional MRI technique, require a human expert to perform segmentation manually. The manual segmentation is error-prone and time-consuming ($\sim$ few hours per subject). We use an ensemble of modified U-Nets to automate this segmentation task. We benchmark our model against a human expert test-retest segmentation and conclude that our model is superior for Patellar and Tibial cartilage using dice score as the comparison metric. In the end, we do a perturbation analysis to understand the sensitivity of our model to the different components of our input. We also provide confidence maps for the predictions so that radiologists can tweak the model predictions as required. The model has been deployed in practice. In conclusion, cartilage segmentation on DW-MRI images with modified U-Nets achieves accuracy that outperforms the human segmenter. Code is available at https://github.com/aakashrkaku/knee-cartilage-segmentation
DARTS: DenseUnet-based Automatic Rapid Tool for brain Segmentation
Nov 14, 2019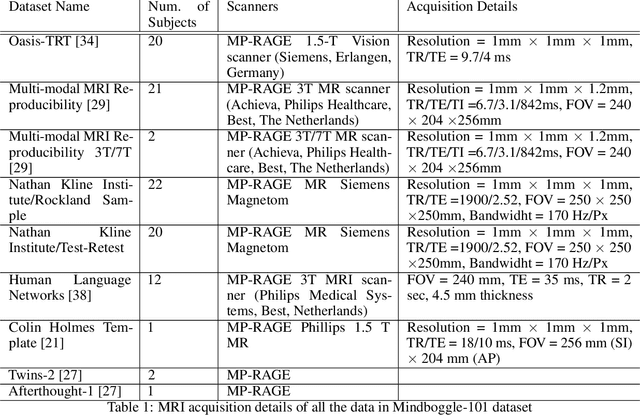
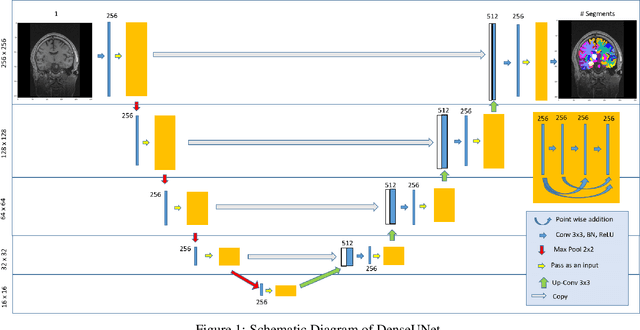
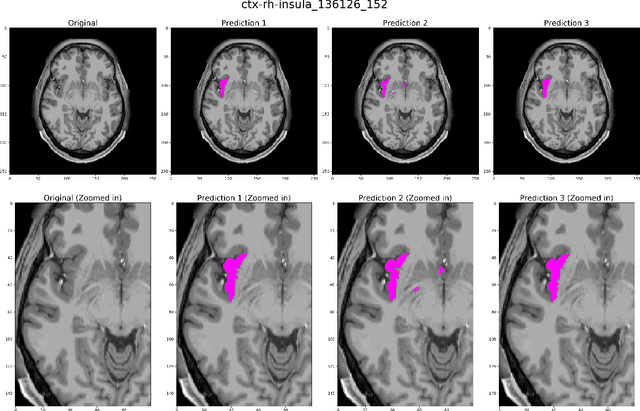
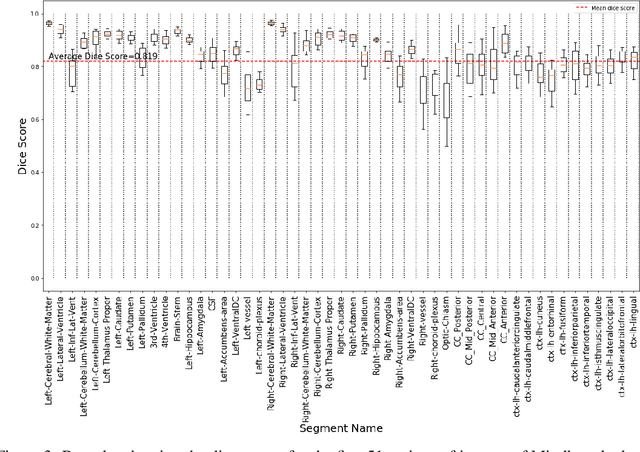
Abstract:Quantitative, volumetric analysis of Magnetic Resonance Imaging (MRI) is a fundamental way researchers study the brain in a host of neurological conditions including normal maturation and aging. Despite the availability of open-source brain segmentation software, widespread clinical adoption of volumetric analysis has been hindered due to processing times and reliance on manual corrections. Here, we extend the use of deep learning models from proof-of-concept, as previously reported, to present a comprehensive segmentation of cortical and deep gray matter brain structures matching the standard regions of aseg+aparc included in the commonly used open-source tool, Freesurfer. The work presented here provides a real-life, rapid deep learning-based brain segmentation tool to enable clinical translation as well as research application of quantitative brain segmentation. The advantages of the presented tool include short (~1 minute) processing time and improved segmentation quality. This is the first study to perform quick and accurate segmentation of 102 brain regions based on the surface-based protocol (DMK protocol), widely used by experts in the field. This is also the first work to include an expert reader study to assess the quality of the segmentation obtained using a deep-learning-based model. We show the superior performance of our deep-learning-based models over the traditional segmentation tool, Freesurfer. We refer to the proposed deep learning-based tool as DARTS (DenseUnet-based Automatic Rapid Tool for brain Segmentation). Our tool and trained models are available at https://github.com/NYUMedML/DARTS
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge