Celia M. T. Greenwood
Varying-Coefficient Mixture of Experts Model
Jan 05, 2026Abstract:Mixture-of-Experts (MoE) is a flexible framework that combines multiple specialized submodels (``experts''), by assigning covariate-dependent weights (``gating functions'') to each expert, and have been commonly used for analyzing heterogeneous data. Existing statistical MoE formulations typically assume constant coefficients, for covariate effects within the expert or gating models, which can be inadequate for longitudinal, spatial, or other dynamic settings where covariate influences and latent subpopulation structure evolve across a known dimension. We propose a Varying-Coefficient Mixture of Experts (VCMoE) model that allows all coefficient effects in both the gating functions and expert models to vary along an indexing variable. We establish identifiability and consistency of the proposed model, and develop an estimation procedure, label-consistent EM algorithm, for both fully functional and hybrid specifications, along with the corresponding asymptotic distributions of the resulting estimators. For inference, simultaneous confidence bands are constructed using both asymptotic theory for the maximum discrepancy between the estimated functional coefficients and their true counterparts, and with bootstrap methods. In addition, a generalized likelihood ratio test is developed to examine whether a coefficient function is genuinely varying across the index variable. Simulation studies demonstrate good finite-sample performance, with acceptable bias and satisfactory coverage rates. We illustrate the proposed VCMoE model using a dataset of single nucleus gene expression in embryonic mice to characterize the temporal dynamics of the associations between the expression levels of genes Satb2 and Bcl11b across two latent cell subpopulations of neurons, yielding results that are consistent with prior findings.
fastHDMI: Fast Mutual Information Estimation for High-Dimensional Data
Oct 14, 2024



Abstract:In this paper, we introduce fastHDMI, a Python package designed for efficient variable screening in high-dimensional datasets, particularly neuroimaging data. This work pioneers the application of three mutual information estimation methods for neuroimaging variable selection, a novel approach implemented via fastHDMI. These advancements enhance our ability to analyze the complex structures of neuroimaging datasets, providing improved tools for variable selection in high-dimensional spaces. Using the preprocessed ABIDE dataset, we evaluate the performance of these methods through extensive simulations. The tests cover a range of conditions, including linear and nonlinear associations, as well as continuous and binary outcomes. Our results highlight the superiority of the FFTKDE-based mutual information estimation for feature screening in continuous nonlinear outcomes, while binning-based methods outperform others for binary outcomes with nonlinear probability preimages. For linear simulations, both Pearson correlation and FFTKDE-based methods show comparable performance for continuous outcomes, while Pearson excels in binary outcomes with linear probability preimages. A comprehensive case study using the ABIDE dataset further demonstrates fastHDMI's practical utility, showcasing the predictive power of models built from variables selected using our screening techniques. This research affirms the computational efficiency and methodological strength of fastHDMI, significantly enriching the toolkit available for neuroimaging analysis.
An algorithm-based multiple detection influence measure for high dimensional regression using expectile
May 26, 2021


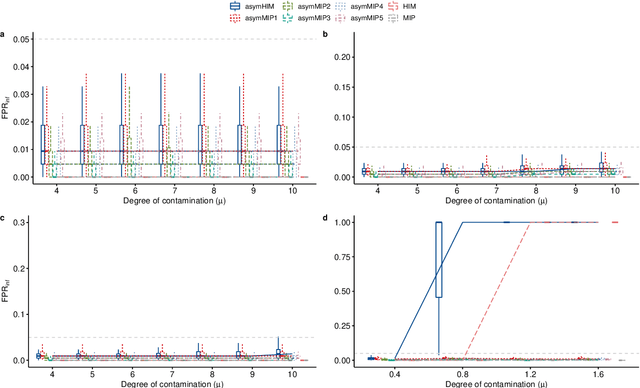
Abstract:The identification of influential observations is an important part of data analysis that can prevent erroneous conclusions drawn from biased estimators. However, in high dimensional data, this identification is challenging. Classical and recently-developed methods often perform poorly when there are multiple influential observations in the same dataset. In particular, current methods can fail when there is masking several influential observations with similar characteristics, or swamping when the influential observations are near the boundary of the space spanned by well-behaved observations. Therefore, we propose an algorithm-based, multi-step, multiple detection procedure to identify influential observations that addresses current limitations. Our three-step algorithm to identify and capture undesirable variability in the data, $\asymMIP,$ is based on two complementary statistics, inspired by asymmetric correlations, and built on expectiles. Simulations demonstrate higher detection power than competing methods. Use of the resulting asymptotic distribution leads to detection of influential observations without the need for computationally demanding procedures such as the bootstrap. The application of our method to the Autism Brain Imaging Data Exchange neuroimaging dataset resulted in a more balanced and accurate prediction of brain maturity based on cortical thickness. See our GitHub for a free R package that implements our algorithm: \texttt{asymMIP} (\url{github.com/AmBarry/hidetify}).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge