Carsten Wiuf
A reaction network scheme which implements inference and learning for Hidden Markov Models
Jun 22, 2019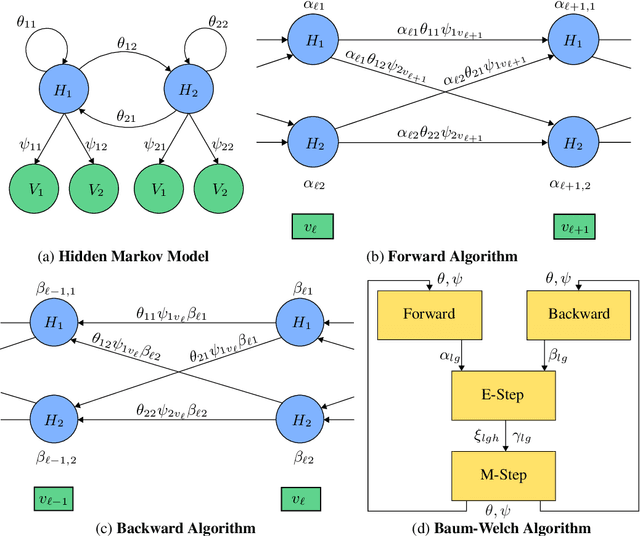


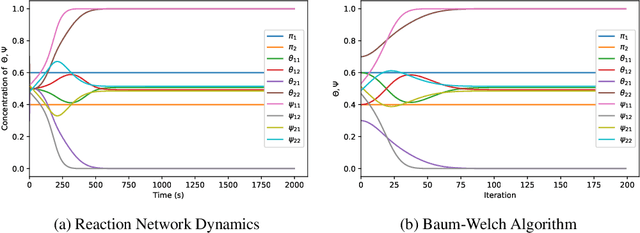
Abstract:With a view towards molecular communication systems and molecular multi-agent systems, we propose the Chemical Baum-Welch Algorithm, a novel reaction network scheme that learns parameters for Hidden Markov Models (HMMs). Each reaction in our scheme changes only one molecule of one species to one molecule of another. The reverse change is also accessible but via a different set of enzymes, in a design reminiscent of futile cycles in biochemical pathways. We show that every fixed point of the Baum-Welch algorithm for HMMs is a fixed point of our reaction network scheme, and every positive fixed point of our scheme is a fixed point of the Baum-Welch algorithm. We prove that the "Expectation" step and the "Maximization" step of our reaction network separately converge exponentially fast. We simulate mass-action kinetics for our network on an example sequence, and show that it learns the same parameters for the HMM as the Baum-Welch algorithm.
Bounded Coordinate-Descent for Biological Sequence Classification in High Dimensional Predictor Space
Aug 03, 2010
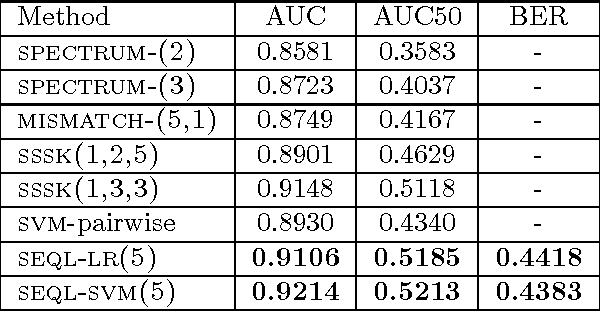


Abstract:We present a framework for discriminative sequence classification where the learner works directly in the high dimensional predictor space of all subsequences in the training set. This is possible by employing a new coordinate-descent algorithm coupled with bounding the magnitude of the gradient for selecting discriminative subsequences fast. We characterize the loss functions for which our generic learning algorithm can be applied and present concrete implementations for logistic regression (binomial log-likelihood loss) and support vector machines (squared hinge loss). Application of our algorithm to protein remote homology detection and remote fold recognition results in performance comparable to that of state-of-the-art methods (e.g., kernel support vector machines). Unlike state-of-the-art classifiers, the resulting classification models are simply lists of weighted discriminative subsequences and can thus be interpreted and related to the biological problem.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge