Busra Celikkaya
Towards User Friendly Medication Mapping Using Entity-Boosted Two-Tower Neural Network
Jun 17, 2020
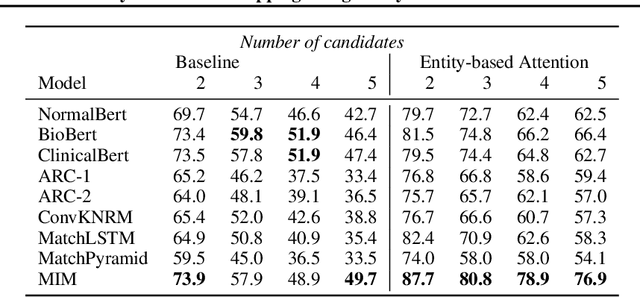
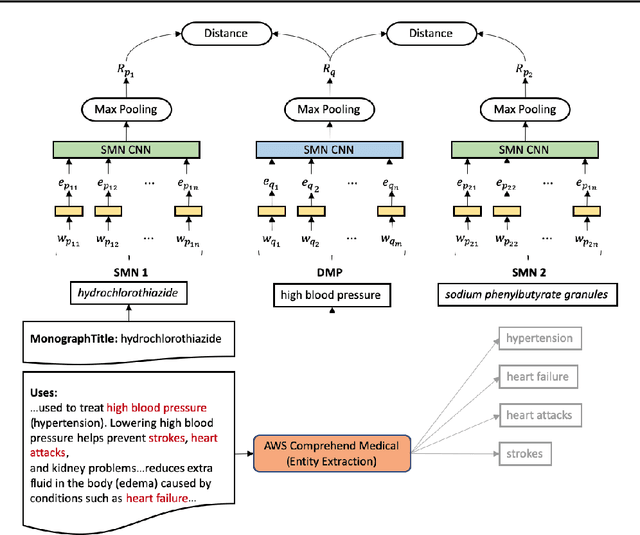

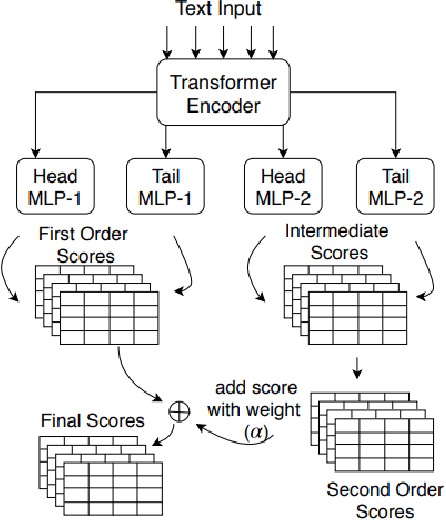
Abstract:Recent advancements in medical entity linking have been applied in the area of scientific literature and social media data. However, with the adoption of telemedicine and conversational agents such as Alexa in healthcare settings, medical name inference has become an important task. Medication name inference is the task of mapping user friendly medication names from a free-form text to a concept in a normalized medication list. This is challenging due to the differences in the use of medical terminology from health care professionals and user conversations coming from the lay public. We begin with mapping descriptive medication phrases (DMP) to standard medication names (SMN). Given the prescriptions of each patient, we want to provide them with the flexibility of referring to the medication in their preferred ways. We approach this as a ranking problem which maps SMN to DMP by ordering the list of medications in the patient's prescription list obtained from pharmacies. Furthermore, we leveraged the output of intermediate layers and performed medication clustering. We present the Medication Inference Model (MIM) achieving state-of-the-art results. By incorporating medical entities based attention, we have obtained further improvement for ranking models.
LATTE: Latent Type Modeling for Biomedical Entity Linking
Nov 21, 2019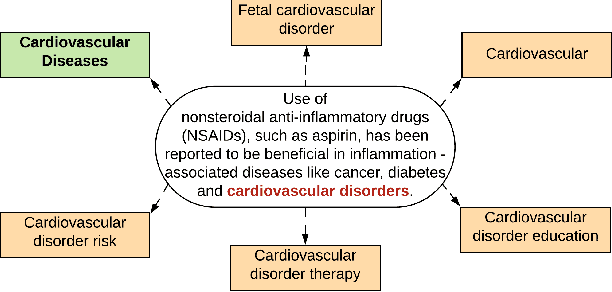
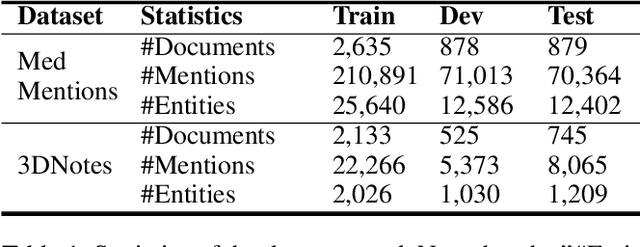
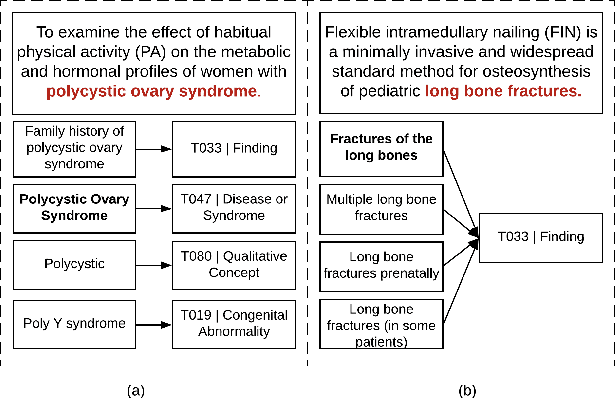
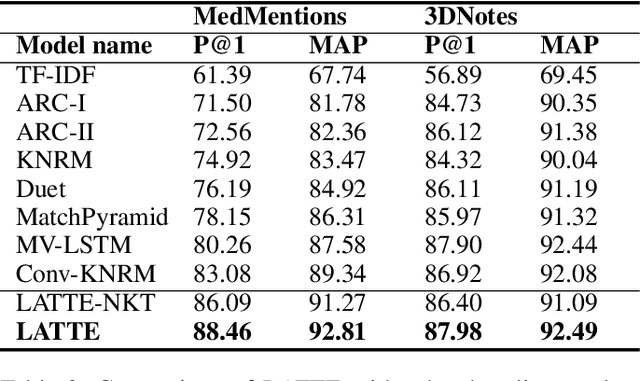
Abstract:Entity linking is the task of linking mentions of named entities in natural language text, to entities in a curated knowledge-base. This is of significant importance in the biomedical domain, where it could be used to semantically annotate a large volume of clinical records and biomedical literature, to standardized concepts described in an ontology such as Unified Medical Language System (UMLS). We observe that with precise type information, entity disambiguation becomes a straightforward task. However, fine-grained type information is usually not available in biomedical domain. Thus, we propose LATTE, a LATent Type Entity Linking model, that improves entity linking by modeling the latent fine-grained type information about mentions and entities. Unlike previous methods that perform entity linking directly between the mentions and the entities, LATTE jointly does entity disambiguation, and latent fine-grained type learning, without direct supervision. We evaluate our model on two biomedical datasets: MedMentions, a large scale public dataset annotated with UMLS concepts, and a de-identified corpus of dictated doctor's notes that has been annotated with ICD concepts. Extensive experimental evaluation shows our model achieves significant performance improvements over several state-of-the-art techniques.
Comprehend Medical: a Named Entity Recognition and Relationship Extraction Web Service
Oct 15, 2019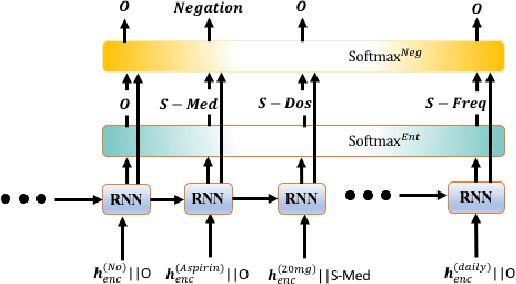
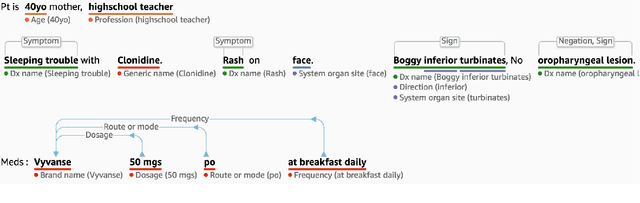
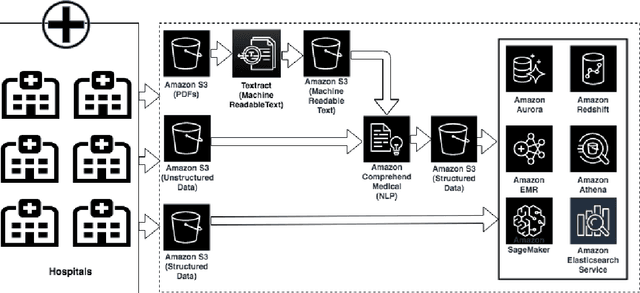
Abstract:Comprehend Medical is a stateless and Health Insurance Portability and Accountability Act (HIPAA) eligible Named Entity Recognition (NER) and Relationship Extraction (RE) service launched under Amazon Web Services (AWS) trained using state-of-the-art deep learning models. Contrary to many existing open source tools, Comprehend Medical is scalable and does not require steep learning curve, dependencies, pipeline configurations, or installations. Currently, Comprehend Medical performs NER in five medical categories: Anatomy, Medical Condition, Medications, Protected Health Information (PHI) and Treatment, Test and Procedure (TTP). Additionally, the service provides relationship extraction for the detected entities as well as contextual information such as negation and temporality in the form of traits. Comprehend Medical provides two Application Programming Interfaces (API): 1) the NERe API which returns all the extracted named entities, their traits and the relationships between them and 2) the PHId API which returns just the protected health information contained in the text. Furthermore, Comprehend Medical is accessible through AWS Console, Java and Python Software Development Kit (SDK), making it easier for non-developers and developers to use.
Dynamic Transfer Learning for Named Entity Recognition
Jan 18, 2019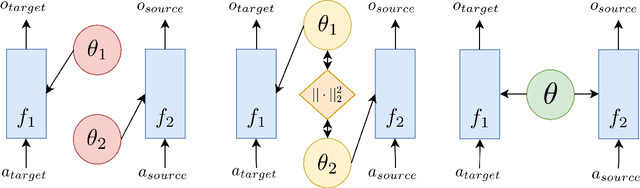
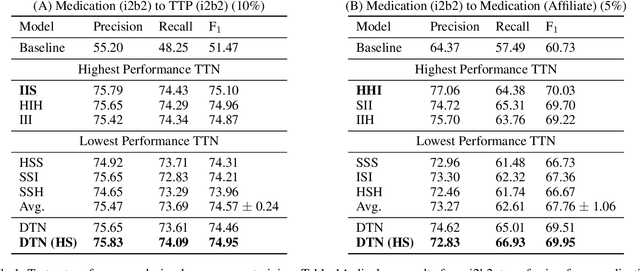
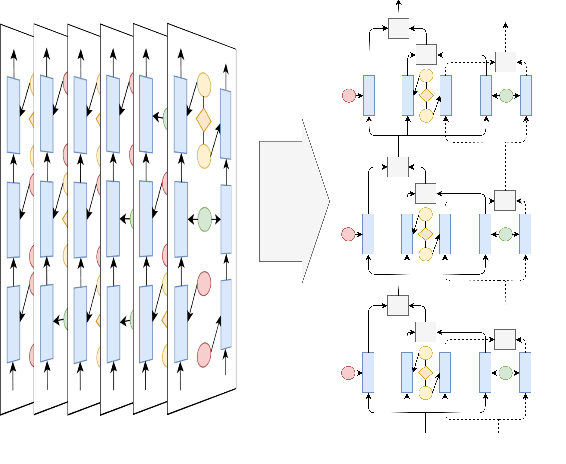
Abstract:State-of-the-art named entity recognition (NER) systems have been improving continuously using neural architectures over the past several years. However, many tasks including NER require large sets of annotated data to achieve such performance. In particular, we focus on NER from clinical notes, which is one of the most fundamental and critical problems for medical text analysis. Our work centers on effectively adapting these neural architectures towards low-resource settings using parameter transfer methods. We complement a standard hierarchical NER model with a general transfer learning framework consisting of parameter sharing between the source and target tasks, and showcase scores significantly above the baseline architecture. These sharing schemes require an exponential search over tied parameter sets to generate an optimal configuration. To mitigate the problem of exhaustively searching for model optimization, we propose the Dynamic Transfer Networks (DTN), a gated architecture which learns the appropriate parameter sharing scheme between source and target datasets. DTN achieves the improvements of the optimized transfer learning framework with just a single training setting, effectively removing the need for exponential search.
End-to-end Joint Entity Extraction and Negation Detection for Clinical Text
Jan 18, 2019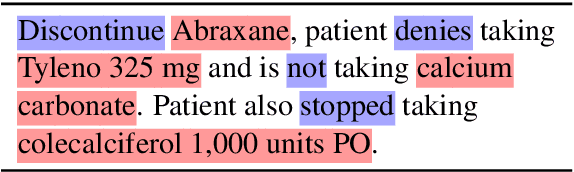
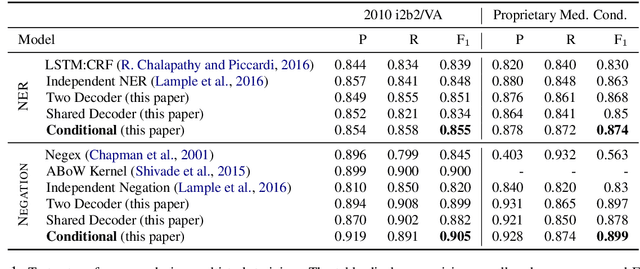
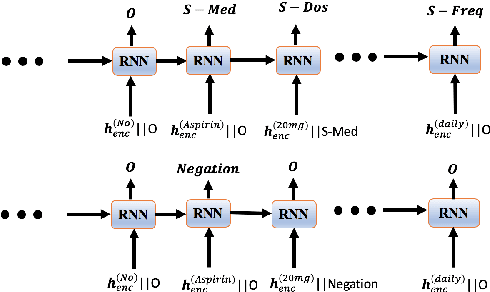
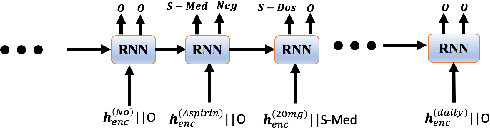
Abstract:Negative medical findings are prevalent in clinical reports, yet discriminating them from positive findings remains a challenging task for information extraction. Most of the existing systems treat this task as a pipeline of two separate tasks, i.e., named entity recognition (NER) and rule-based negation detection. We consider this as a multi-task problem and present a novel end-to-end neural model to jointly extract entities and negations. We extend a standard hierarchical encoder-decoder NER model and first adopt a shared encoder followed by separate decoders for the two tasks. This architecture performs considerably better than the previous rule-based and machine learning-based systems. To overcome the problem of increased parameter size especially for low-resource settings, we propose the \textit{Conditional Softmax Shared Decoder} architecture which achieves state-of-art results for NER and negation detection on the 2010 i2b2/VA challenge dataset and a proprietary de-identified clinical dataset.
Improving Hospital Mortality Prediction with Medical Named Entities and Multimodal Learning
Dec 04, 2018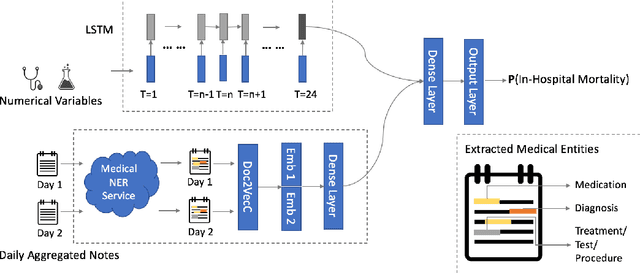
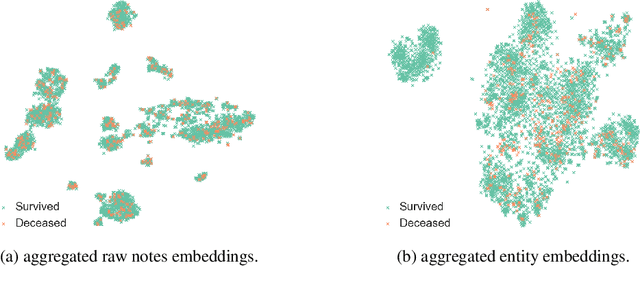
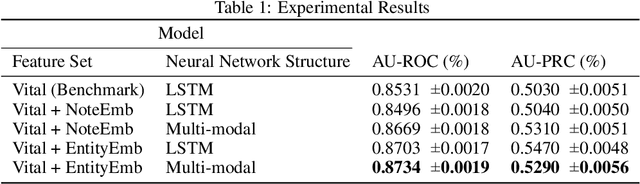

Abstract:Clinical text provides essential information to estimate the acuity of a patient during hospital stays in addition to structured clinical data. In this study, we explore how clinical text can complement a clinical predictive learning task. We leverage an internal medical natural language processing service to perform named entity extraction and negation detection on clinical notes and compose selected entities into a new text corpus to train document representations. We then propose a multimodal neural network to jointly train time series signals and unstructured clinical text representations to predict the in-hospital mortality risk for ICU patients. Our model outperforms the benchmark by 2% AUC.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge