Boudewijn Lelieveldt
Manifold-Preserving Superpixel Hierarchies and Embeddings for the Exploration of High-Dimensional Images
Feb 27, 2026Abstract:High-dimensional images, or images with a high-dimensional attribute vector per pixel, are commonly explored with coordinated views of a low-dimensional embedding of the attribute space and a conventional image representation. Nowadays, such images can easily contain several million pixels. For such large datasets, hierarchical embedding techniques are better suited to represent the high-dimensional attribute space than flat dimensionality reduction methods. However, available hierarchical dimensionality reduction methods construct the hierarchy purely based on the attribute information and ignore the spatial layout of pixels in the images. This impedes the exploration of regions of interest in the image space, since there is no congruence between a region of interest in image space and the associated attribute abstractions in the hierarchy. In this paper, we present a superpixel hierarchy for high-dimensional images that takes the high-dimensional attribute manifold into account during construction. Through this, our method enables consistent exploration of high-dimensional images in both image and attribute space. We show the effectiveness of this new image-guided hierarchy in the context of embedding exploration by comparing it with classical hierarchical embedding-based image exploration in two use cases.
Incorporating Texture Information into Dimensionality Reduction for High-Dimensional Images
Mar 02, 2022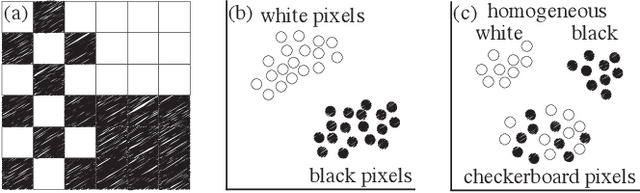
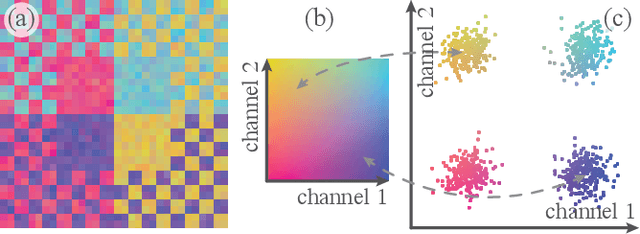
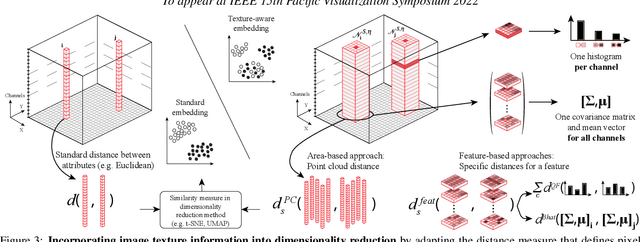
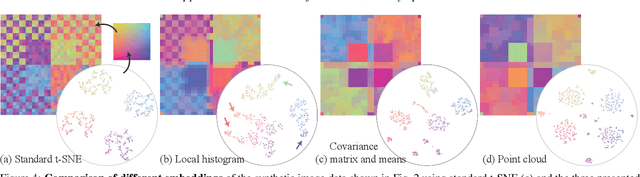
Abstract:High-dimensional imaging is becoming increasingly relevant in many fields from astronomy and cultural heritage to systems biology. Visual exploration of such high-dimensional data is commonly facilitated by dimensionality reduction. However, common dimensionality reduction methods do not include spatial information present in images, such as local texture features, into the construction of low-dimensional embeddings. Consequently, exploration of such data is typically split into a step focusing on the attribute space followed by a step focusing on spatial information, or vice versa. In this paper, we present a method for incorporating spatial neighborhood information into distance-based dimensionality reduction methods, such as t-Distributed Stochastic Neighbor Embedding (t-SNE). We achieve this by modifying the distance measure between high-dimensional attribute vectors associated with each pixel such that it takes the pixel's spatial neighborhood into account. Based on a classification of different methods for comparing image patches, we explore a number of different approaches. We compare these approaches from a theoretical and experimental point of view. Finally, we illustrate the value of the proposed methods by qualitative and quantitative evaluation on synthetic data and two real-world use cases.
Hierarchical stochastic neighbor embedding as a tool for visualizing the encoding capability of magnetic resonance fingerprinting dictionaries
Oct 07, 2019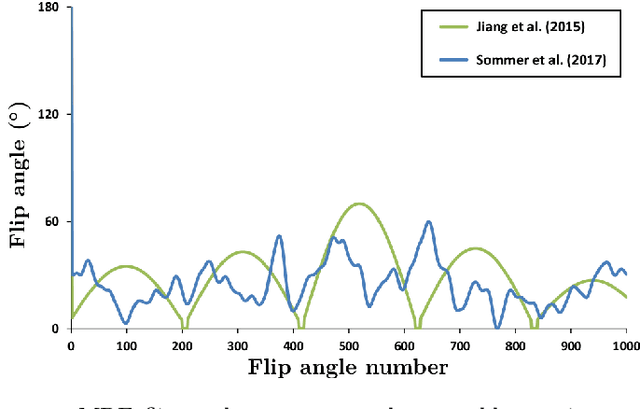
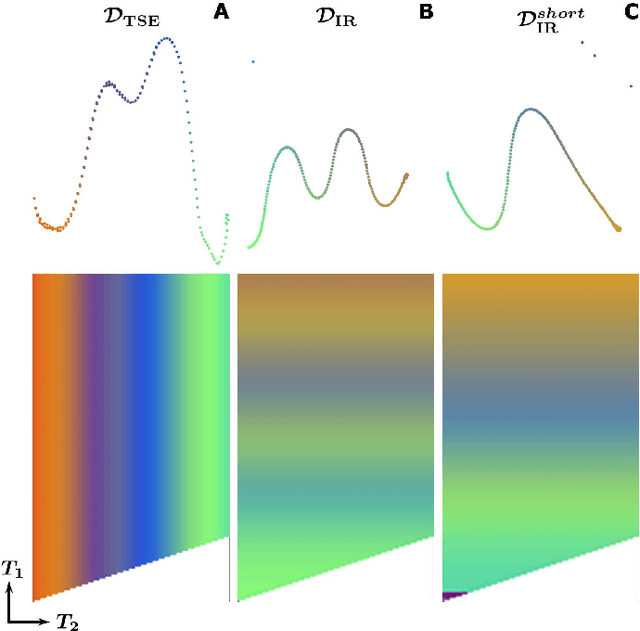
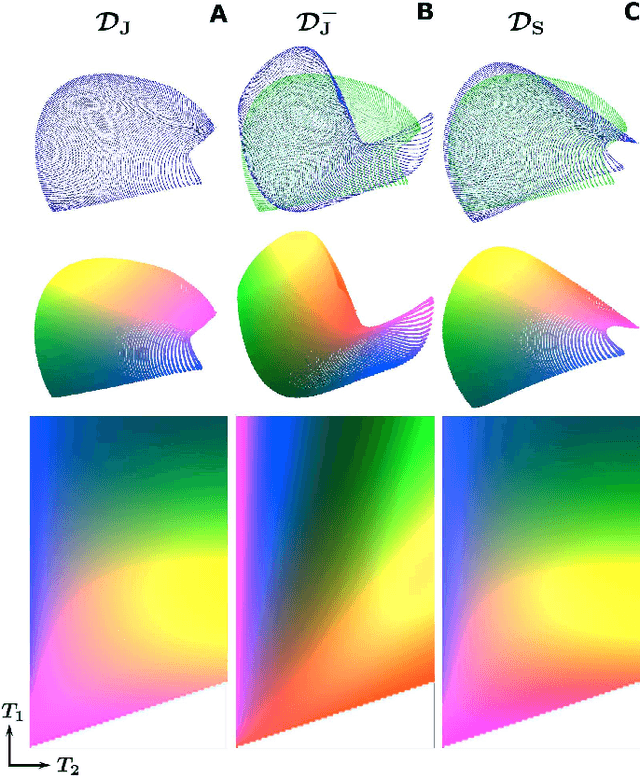
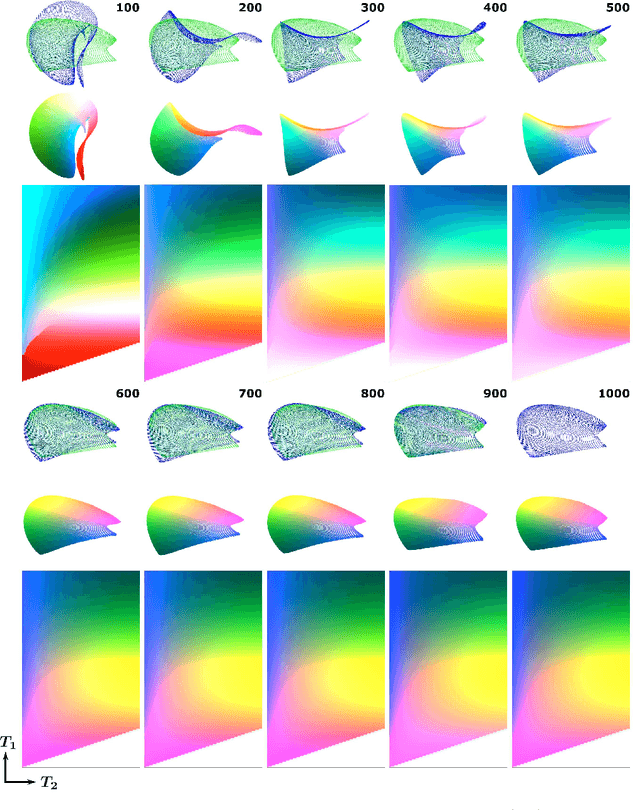
Abstract:In Magnetic Resonance Fingerprinting (MRF) the quality of the estimated parameter maps depends on the encoding capability of the variable flip angle train. In this work we show how the dimensionality reduction technique Hierarchical Stochastic Neighbor Embedding (HSNE) can be used to obtain insight into the encoding capability of different MRF sequences. Embedding high-dimensional MRF dictionaries into a lower-dimensional space and visualizing them with colors, being a surrogate for location in low-dimensional space, provides a comprehensive overview of particular dictionaries and, in addition, enables comparison of different sequences. Dictionaries for various sequences and sequence lengths were compared to each other, and the effect of transmit field variations on the encoding capability was assessed. Clear differences in encoding capability were observed between different sequences, and HSNE results accurately reflect those obtained from an MRF matching simulation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge