Bino Varghese
Longitudinal Ensemble Integration for sequential classification with multimodal data
Nov 08, 2024



Abstract:Effectively modeling multimodal longitudinal data is a pressing need in various application areas, especially biomedicine. Despite this, few approaches exist in the literature for this problem, with most not adequately taking into account the multimodality of the data. In this study, we developed multiple configurations of a novel multimodal and longitudinal learning framework, Longitudinal Ensemble Integration (LEI), for sequential classification. We evaluated LEI's performance, and compared it against existing approaches, for the early detection of dementia, which is among the most studied multimodal sequential classification tasks. LEI outperformed these approaches due to its use of intermediate base predictions arising from the individual data modalities, which enabled their better integration over time. LEI's design also enabled the identification of features that were consistently important across time for the effective prediction of dementia-related diagnoses. Overall, our work demonstrates the potential of LEI for sequential classification from longitudinal multimodal data.
Benchmarking features from different radiomics toolkits / toolboxes using Image Biomarkers Standardization Initiative
Jun 23, 2020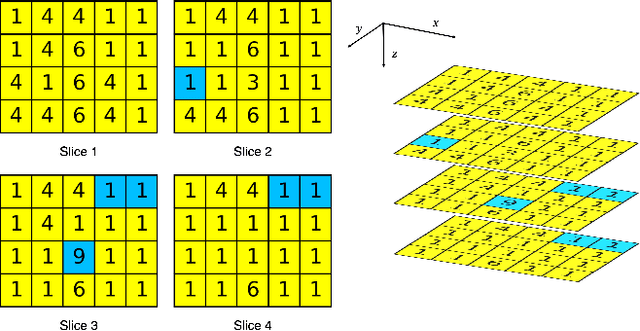
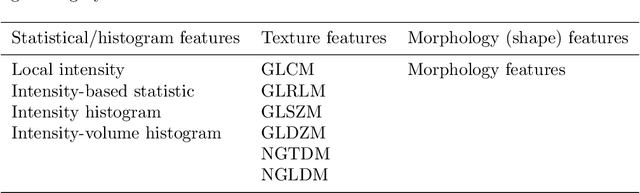
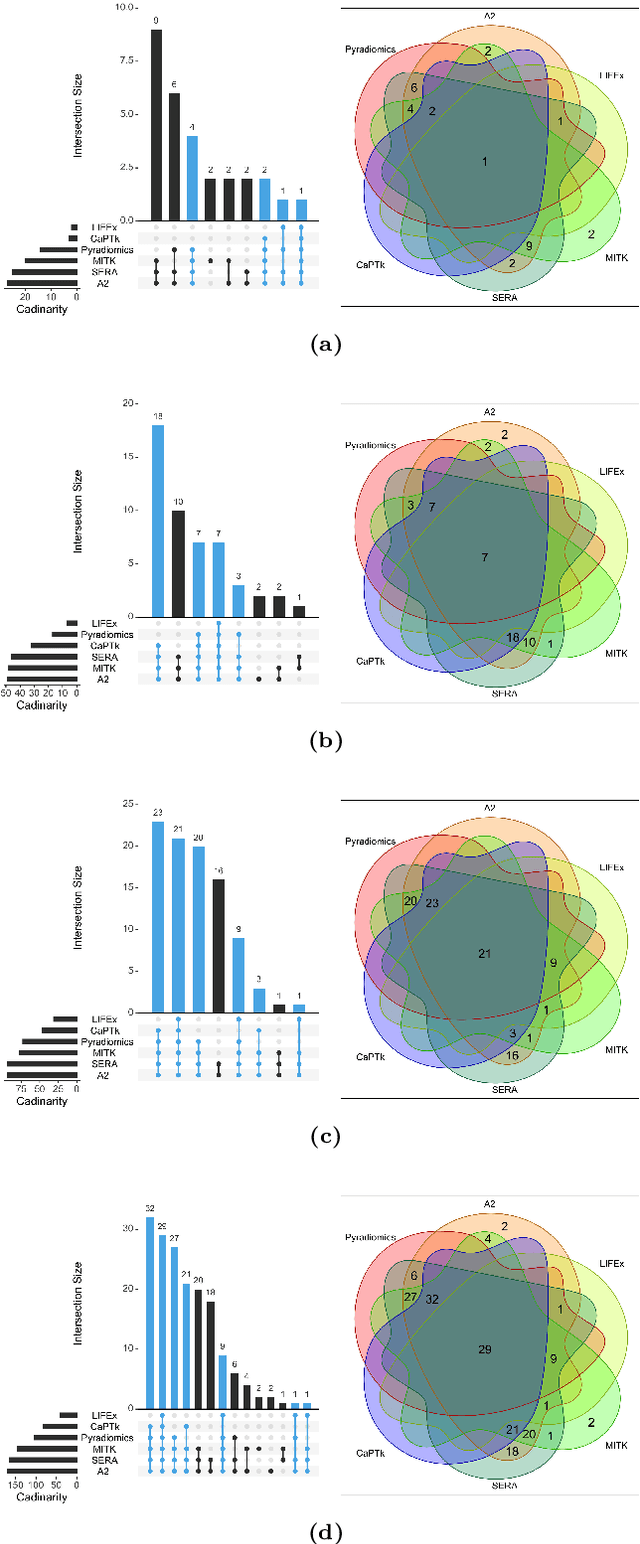
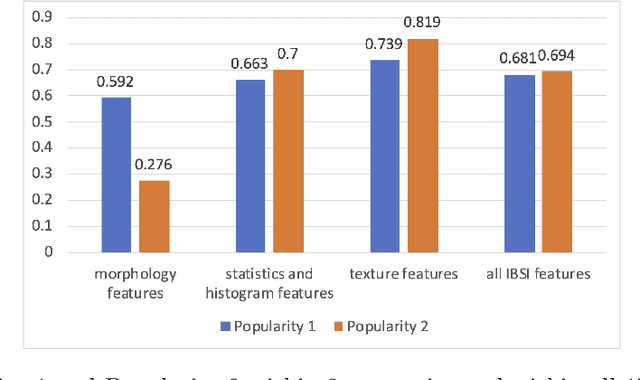
Abstract:There is no consensus regarding the radiomic feature terminology, the underlying mathematics, or their implementation. This creates a scenario where features extracted using different toolboxes could not be used to build or validate the same model leading to a non-generalization of radiomic results. In this study, the image biomarker standardization initiative (IBSI) established phantom and benchmark values were used to compare the variation of the radiomic features while using 6 publicly available software programs and 1 in-house radiomics pipeline. All IBSI-standardized features (11 classes, 173 in total) were extracted. The relative differences between the extracted feature values from the different software and the IBSI benchmark values were calculated to measure the inter-software agreement. To better understand the variations, features are further grouped into 3 categories according to their properties: 1) morphology, 2) statistic/histogram and 3)texture features. While a good agreement was observed for a majority of radiomics features across the various programs, relatively poor agreement was observed for morphology features. Significant differences were also found in programs that use different gray level discretization approaches. Since these programs do not include all IBSI features, the level of quantitative assessment for each category was analyzed using Venn and the UpSet diagrams and also quantified using two ad hoc metrics. Morphology features earns lowest scores for both metrics, indicating that morphological features are not consistently evaluated among software programs. We conclude that radiomic features calculated using different software programs may not be identical and reliable. Further studies are needed to standardize the workflow of radiomic feature extraction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge