Bernhard Strasser
Athinoula A. Martinos Center for Biomedical Imaging, Department of Radiology, Massachusetts General Hospital, Harvard Medical School, Boston, High-Field MR Center, Department of Biomedical Imaging and Image-guided Therapy, Medical University of Vienna, Vienna, Austria
Deep-ER: Deep Learning ECCENTRIC Reconstruction for fast high-resolution neurometabolic imaging
Sep 26, 2024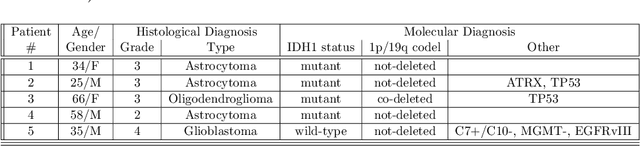
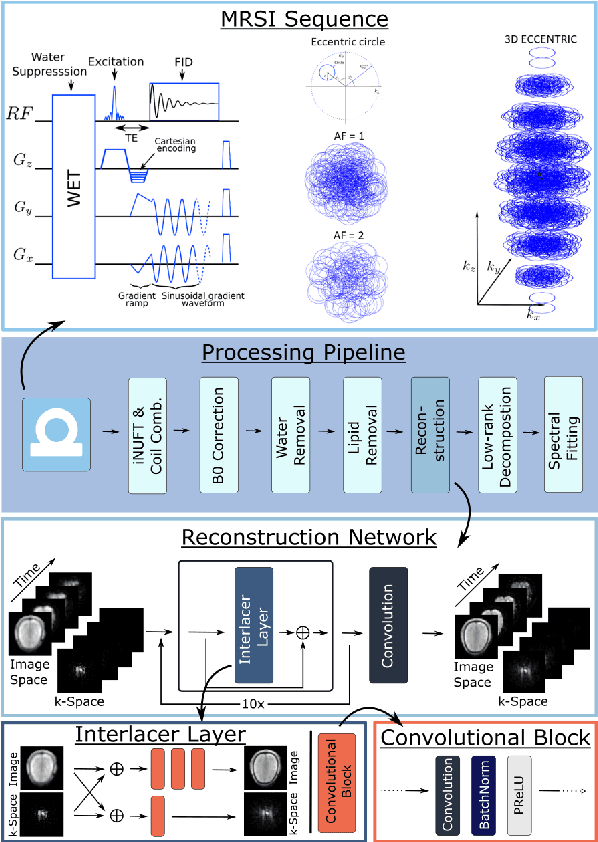
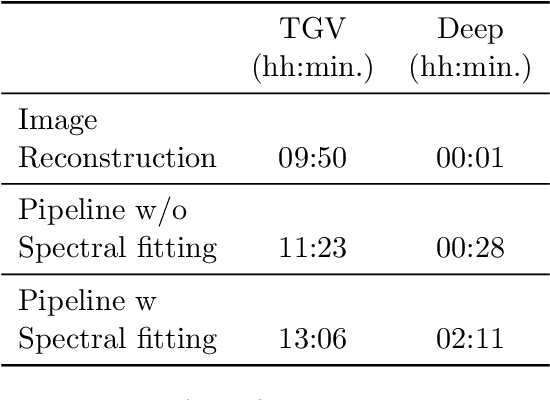

Abstract:Introduction: Altered neurometabolism is an important pathological mechanism in many neurological diseases and brain cancer, which can be mapped non-invasively by Magnetic Resonance Spectroscopic Imaging (MRSI). Advanced MRSI using non-cartesian compressed-sense acquisition enables fast high-resolution metabolic imaging but has lengthy reconstruction times that limits throughput and needs expert user interaction. Here, we present a robust and efficient Deep Learning reconstruction to obtain high-quality metabolic maps. Methods: Fast high-resolution whole-brain metabolic imaging was performed at 3.4 mm$^3$ isotropic resolution with acquisition times between 4:11-9:21 min:s using ECCENTRIC pulse sequence on a 7T MRI scanner. Data were acquired in a high-resolution phantom and 27 human participants, including 22 healthy volunteers and 5 glioma patients. A deep neural network using recurring interlaced convolutional layers with joint dual-space feature representation was developed for deep learning ECCENTRIC reconstruction (Deep-ER). 21 subjects were used for training and 6 subjects for testing. Deep-ER performance was compared to conventional iterative Total Generalized Variation reconstruction using image and spectral quality metrics. Results: Deep-ER demonstrated 600-fold faster reconstruction than conventional methods, providing improved spatial-spectral quality and metabolite quantification with 12%-45% (P<0.05) higher signal-to-noise and 8%-50% (P<0.05) smaller Cramer-Rao lower bounds. Metabolic images clearly visualize glioma tumor heterogeneity and boundary. Conclusion: Deep-ER provides efficient and robust reconstruction for sparse-sampled MRSI. The accelerated acquisition-reconstruction MRSI is compatible with high-throughput imaging workflow. It is expected that such improved performance will facilitate basic and clinical MRSI applications.
ECCENTRIC: a fast and unrestrained approach for high-resolution in vivo metabolic imaging at ultra-high field MR
May 23, 2023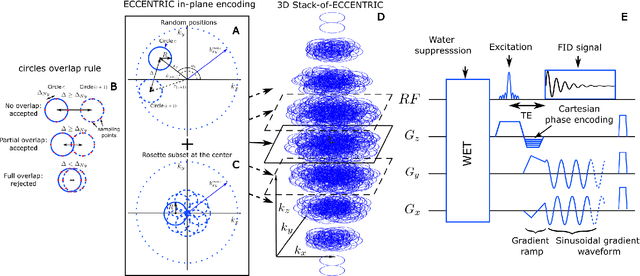
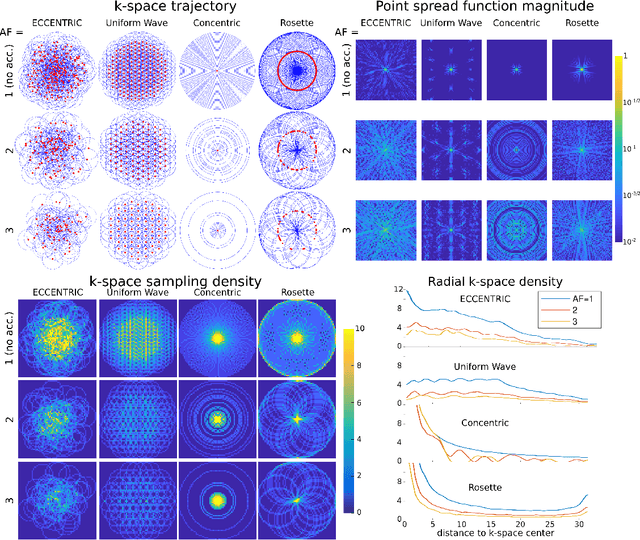
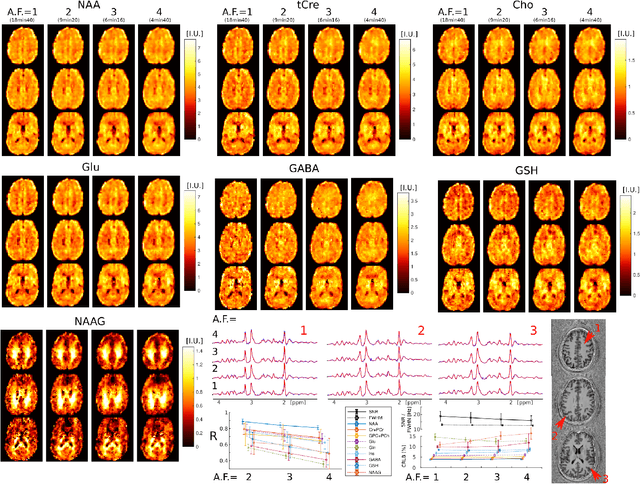
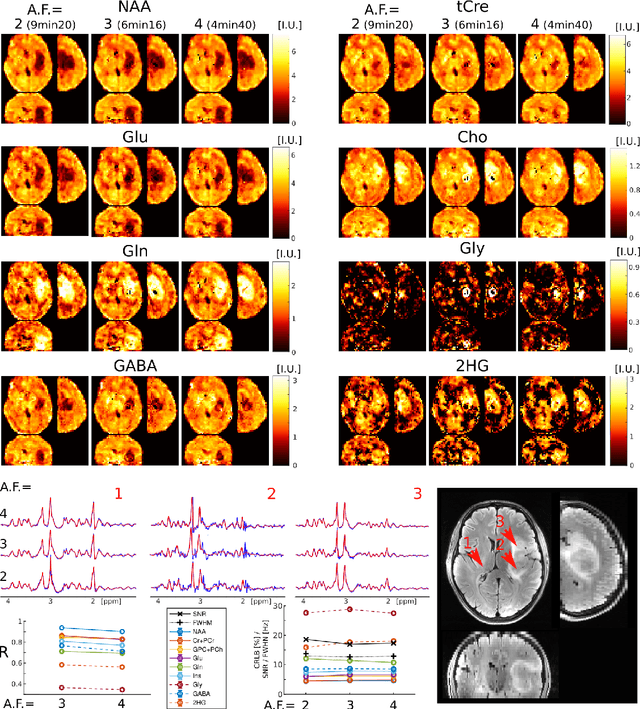
Abstract:A novel method for fast and high-resolution metabolic imaging, called ECcentric Circle ENcoding TRajectorIes for Compressed sensing (ECCENTRIC), has been developed and implemented on 7 Tesla human MRI. ECCENTRIC is a non-Cartesian spatial-spectral encoding method optimized for random undersampling of magnetic resonance spectroscopic imaging (MRSI) at ultra-high field. The approach provides flexible and random (k,t) sampling without temporal interleaving to improve spatial response function and spectral quality. ECCENTRIC needs low gradient amplitudes and slew-rates that reduces electrical, mechanical and thermal stress of the scanner hardware, and is robust to timing imperfection and eddy-current delays. Combined with a model-based low-rank reconstruction, this approach enables simultaneous imaging of up to 14 metabolites over the whole-brain at 2-3mm isotropic resolution in 4-10 minutes with high signal-to-noise ratio. In 20 healthy volunteers and 20 glioma patients ECCENTRIC demonstrated unprecedented mapping of fine structural details of metabolism in healthy brains and an extended metabolic fingerprinting of glioma tumors.
Predicting dynamic, motion-related changes in B0 field in the brain at a 7 T MRI using a subject-specific fine-tuned U-net
Apr 17, 2023Abstract:Subject movement during the magnetic resonance examination is inevitable and causes not only image artefacts but also deteriorates the homogeneity of the main magnetic field (B0), which is a prerequisite for high quality data. Thus, characterization of changes to B0, e.g. induced by patient movement, is important for MR applications that are prone to B0 inhomogeneities. We propose a deep learning based method to predict such changes within the brain from the change of the head position to facilitate retrospective or even real-time correction. A 3D U-net was trained on in vivo brain 7T MRI data. The input consisted of B0 maps and anatomical images at an initial position, and anatomical images at a different head position (obtained by applying a rigid-body transformation on the initial anatomical image). The output consisted of B0 maps at the new head positions. We further fine-tuned the network weights to each subject by measuring a limited number of head positions of the given subject, and trained the U-net with these data. Our approach was compared to established dynamic B0 field mapping via interleaved navigators, which suffer from limited spatial resolution and the need for undesirable sequence modifications. Qualitative and quantitative comparison showed similar performance between an interleaved navigator-equivalent method and proposed method. We therefore conclude that it is feasible to predict B0 maps from rigid subject movement and, when combined with external tracking hardware, this information could be used to improve the quality of magnetic resonance acquisitions without the use of navigators.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge