Avan Suinesiaputra
Neural Implicit Heart Coordinates: 3D cardiac shape reconstruction from sparse segmentations
Dec 23, 2025Abstract:Accurate reconstruction of cardiac anatomy from sparse clinical images remains a major challenge in patient-specific modeling. While neural implicit functions have previously been applied to this task, their application to mapping anatomical consistency across subjects has been limited. In this work, we introduce Neural Implicit Heart Coordinates (NIHCs), a standardized implicit coordinate system, based on universal ventricular coordinates, that provides a common anatomical reference frame for the human heart. Our method predicts NIHCs directly from a limited number of 2D segmentations (sparse acquisition) and subsequently decodes them into dense 3D segmentations and high-resolution meshes at arbitrary output resolution. Trained on a large dataset of 5,000 cardiac meshes, the model achieves high reconstruction accuracy on clinical contours, with mean Euclidean surface errors of 2.51$\pm$0.33 mm in a diseased cohort (n=4549) and 2.3$\pm$0.36 mm in a healthy cohort (n=5576). The NIHC representation enables anatomically coherent reconstruction even under severe slice sparsity and segmentation noise, faithfully recovering complex structures such as the valve planes. Compared with traditional pipelines, inference time is reduced from over 60 s to 5-15 s. These results demonstrate that NIHCs constitute a robust and efficient anatomical representation for patient-specific 3D cardiac reconstruction from minimal input data.
Deep Learning Analysis of Cardiac MRI in Legacy Datasets: Multi-Ethnic Study of Atherosclerosis
Oct 28, 2021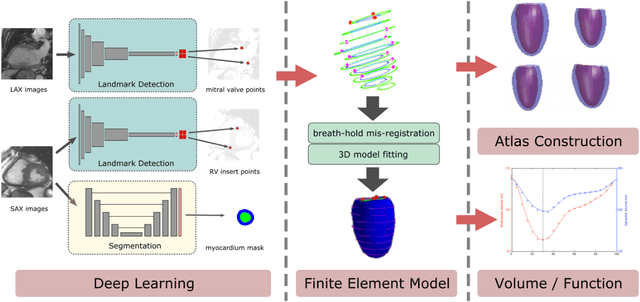
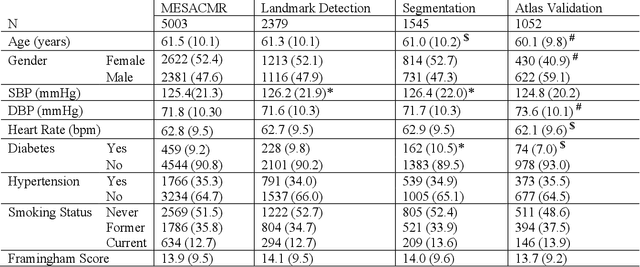
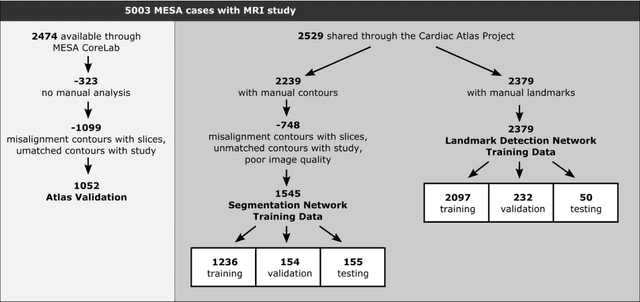
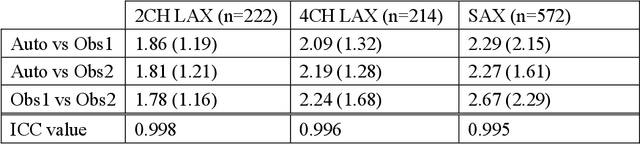
Abstract:The shape and motion of the heart provide essential clues to understanding the mechanisms of cardiovascular disease. With the advent of large-scale cardiac imaging data, statistical atlases become a powerful tool to provide automated and precise quantification of the status of patient-specific heart geometry with respect to reference populations. The Multi-Ethnic Study of Atherosclerosis (MESA), begun in 2000, was the first large cohort study to incorporate cardiovascular MRI in over 5000 participants, and there is now a wealth of follow-up data over 20 years. Building a machine learning based automated analysis is necessary to extract the additional imaging information necessary for expanding original manual analyses. However, machine learning tools trained on MRI datasets with different pulse sequences fail on such legacy datasets. Here, we describe an automated atlas construction pipeline using deep learning methods applied to the legacy cardiac MRI data in MESA. For detection of anatomical cardiac landmark points, a modified VGGNet convolutional neural network architecture was used in conjunction with a transfer learning sequence between two-chamber, four-chamber, and short-axis MRI views. A U-Net architecture was used for detection of the endocardial and epicardial boundaries in short axis images. Both network architectures resulted in good segmentation and landmark detection accuracies compared with inter-observer variations. Statistical relationships with common risk factors were similar between atlases derived from automated vs manual annotations. The automated atlas can be employed in future studies to examine the relationships between cardiac morphology and future events.
Fully Automated Myocardial Strain Estimation from CMR Tagged Images using a Deep Learning Framework in the UK Biobank
Apr 15, 2020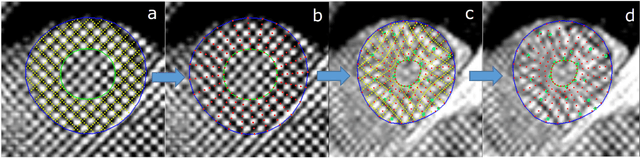
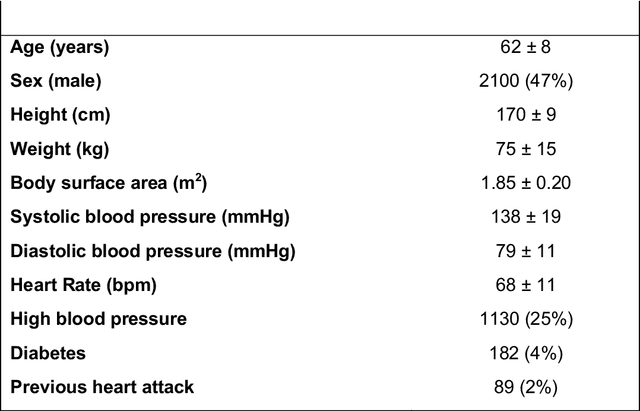

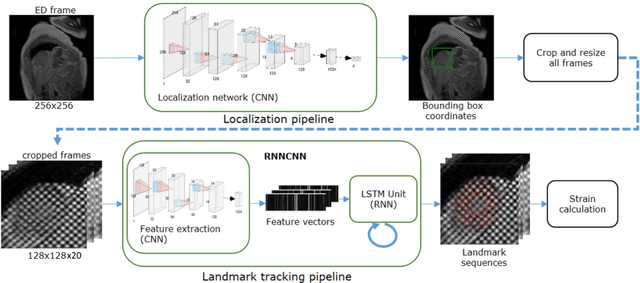
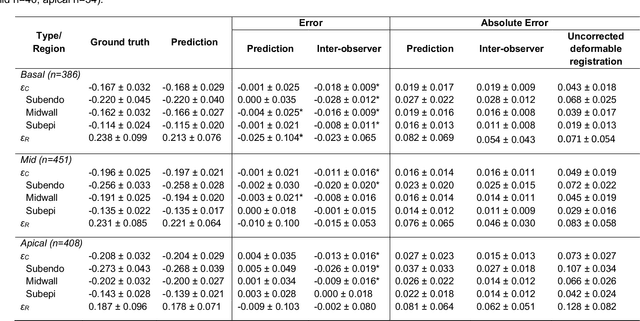
Abstract:Purpose: To demonstrate the feasibility and performance of a fully automated deep learning framework to estimate myocardial strain from short-axis cardiac magnetic resonance tagged images. Methods and Materials: In this retrospective cross-sectional study, 4508 cases from the UK Biobank were split randomly into 3244 training and 812 validation cases, and 452 test cases. Ground truth myocardial landmarks were defined and tracked by manual initialization and correction of deformable image registration using previously validated software with five readers. The fully automatic framework consisted of 1) a convolutional neural network (CNN) for localization, and 2) a combination of a recurrent neural network (RNN) and a CNN to detect and track the myocardial landmarks through the image sequence for each slice. Radial and circumferential strain were then calculated from the motion of the landmarks and averaged on a slice basis. Results: Within the test set, myocardial end-systolic circumferential Green strain errors were -0.001 +/- 0.025, -0.001 +/- 0.021, and 0.004 +/- 0.035 in basal, mid, and apical slices respectively (mean +/- std. dev. of differences between predicted and manual strain). The framework reproduced significant reductions in circumferential strain in diabetics, hypertensives, and participants with previous heart attack. Typical processing time was ~260 frames (~13 slices) per second on an NVIDIA Tesla K40 with 12GB RAM, compared with 6-8 minutes per slice for the manual analysis. Conclusions: The fully automated RNNCNN framework for analysis of myocardial strain enabled unbiased strain evaluation in a high-throughput workflow, with similar ability to distinguish impairment due to diabetes, hypertension, and previous heart attack.
* accepted in Radiology Cardiothoracic Imaging
4DFlowNet: Super-Resolution 4D Flow MRI using Deep Learning and Computational Fluid Dynamics
Apr 15, 2020
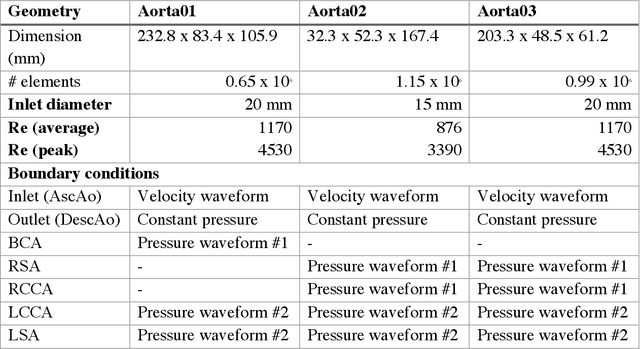

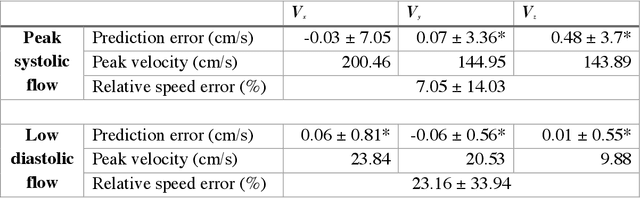
Abstract:4D-flow magnetic resonance imaging (MRI) is an emerging imaging technique where spatiotemporal 3D blood velocity can be captured with full volumetric coverage in a single non-invasive examination. This enables qualitative and quantitative analysis of hemodynamic flow parameters of the heart and great vessels. An increase in the image resolution would provide more accuracy and allow better assessment of the blood flow, especially for patients with abnormal flows. However, this must be balanced with increasing imaging time. The recent success of deep learning in generating super resolution images shows promise for implementation in medical images. We utilized computational fluid dynamics simulations to generate fluid flow simulations and represent them as synthetic 4D flow MRI data. We built our training dataset to mimic actual 4D flow MRI data with its corresponding noise distribution. Our novel 4DFlowNet network was trained on this synthetic 4D flow data and was capable in producing noise-free super resolution 4D flow phase images with upsample factor of 2. We also tested the 4DFlowNet in actual 4D flow MR images of a phantom and normal volunteer data, and demonstrated comparable results with the actual flow rate measurements giving an absolute relative error of 0.6 to 5.8% and 1.1 to 3.8% in the phantom data and normal volunteer data, respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge