Aurelia Bustos
AI Cancer Research Unit Medbravo
PadChest-GR: A Bilingual Chest X-ray Dataset for Grounded Radiology Report Generation
Nov 07, 2024



Abstract:Radiology report generation (RRG) aims to create free-text radiology reports from clinical imaging. Grounded radiology report generation (GRRG) extends RRG by including the localisation of individual findings on the image. Currently, there are no manually annotated chest X-ray (CXR) datasets to train GRRG models. In this work, we present a dataset called PadChest-GR (Grounded-Reporting) derived from PadChest aimed at training GRRG models for CXR images. We curate a public bi-lingual dataset of 4,555 CXR studies with grounded reports (3,099 abnormal and 1,456 normal), each containing complete lists of sentences describing individual present (positive) and absent (negative) findings in English and Spanish. In total, PadChest-GR contains 7,037 positive and 3,422 negative finding sentences. Every positive finding sentence is associated with up to two independent sets of bounding boxes labelled by different readers and has categorical labels for finding type, locations, and progression. To the best of our knowledge, PadChest-GR is the first manually curated dataset designed to train GRRG models for understanding and interpreting radiological images and generated text. By including detailed localization and comprehensive annotations of all clinically relevant findings, it provides a valuable resource for developing and evaluating GRRG models from CXR images. PadChest-GR can be downloaded under request from https://bimcv.cipf.es/bimcv-projects/padchest-gr/
XDEEP-MSI: Explainable Bias-Rejecting Microsatellite Instability Deep Learning System In Colorectal Cancer
Oct 28, 2021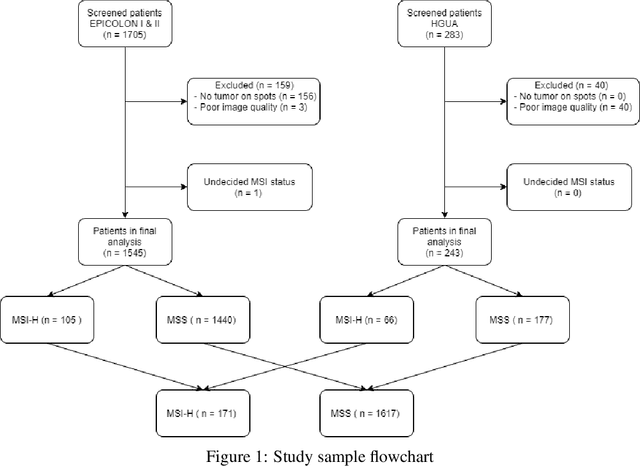
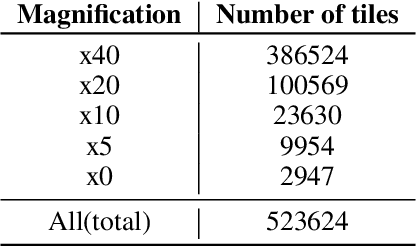

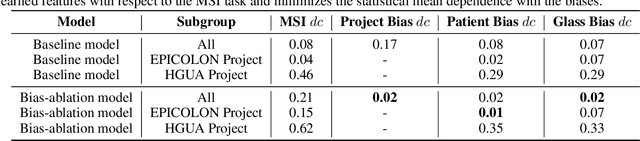
Abstract:We present a system for the prediction of microsatellite instability (MSI) from H&E images of colorectal cancer using deep learning (DL) techniques customized for tissue microarrays (TMAs). The system incorporates an end-to-end image preprocessing module that produces tiles at multiple magnifications in the regions of interest as guided by a tissue classifier module, and a multiple-bias rejecting module. The training and validation TMA samples were obtained from the EPICOLON project and further enriched with samples from a single institution. A systematic study of biases at tile level identified three protected (bias) variables associated with the learned representations of a baseline model: the project of origin of samples, the patient spot and the TMA glass where each spot was placed. A multiple bias rejecting technique based on adversarial training is implemented at the DL architecture so to directly avoid learning the batch effects of those variables. The learned features from the bias-ablated model have maximum discriminative power with respect to the task and minimal statistical mean dependence with the biases. The impact of different magnifications, types of tissues and the model performance at tile vs patient level is analyzed. The AUC at tile level, and including all three selected tissues (tumor epithelium, mucine and lymphocytic regions) and 4 magnifications, was 0.87 +/- 0.03 and increased to 0.9 +/- 0.03 at patient level. To the best of our knowledge, this is the first work that incorporates a multiple bias ablation technique at the DL architecture in digital pathology, and the first using TMAs for the MSI prediction task.
Machine Learning for Real-World Evidence Analysis of COVID-19 Pharmacotherapy
Jul 19, 2021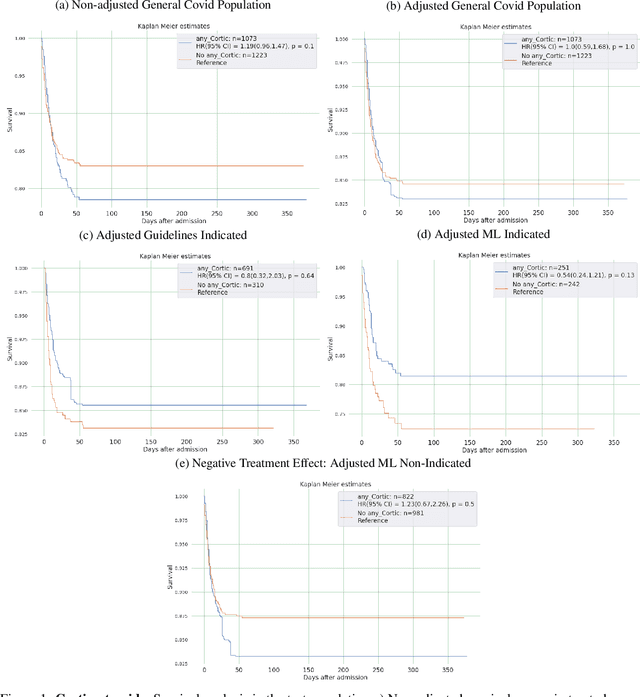
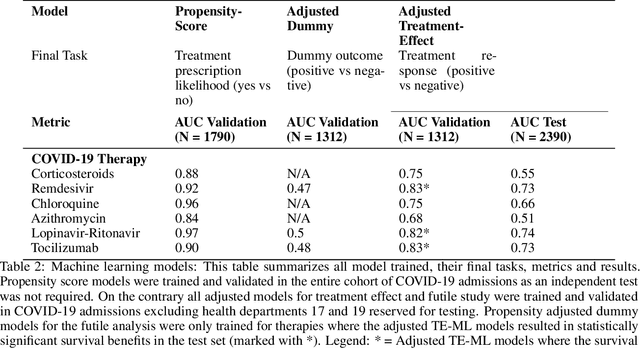
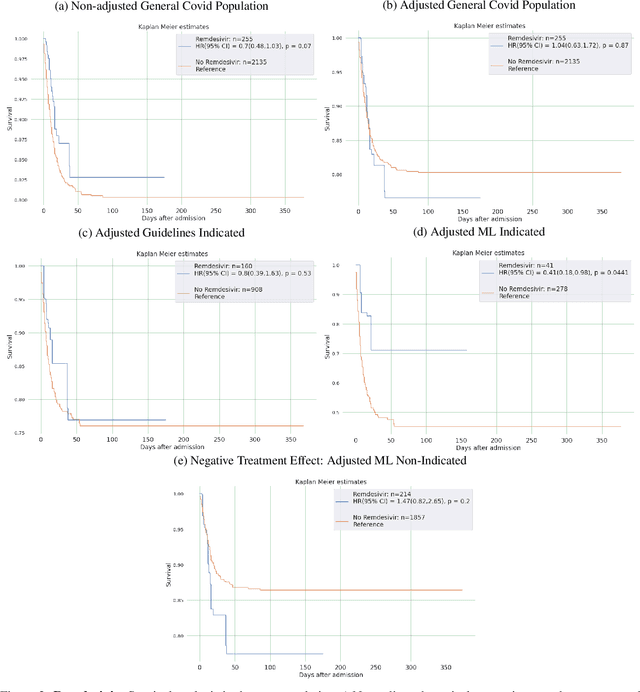

Abstract:Introduction: Real-world data generated from clinical practice can be used to analyze the real-world evidence (RWE) of COVID-19 pharmacotherapy and validate the results of randomized clinical trials (RCTs). Machine learning (ML) methods are being used in RWE and are promising tools for precision-medicine. In this study, ML methods are applied to study the efficacy of therapies on COVID-19 hospital admissions in the Valencian Region in Spain. Methods: 5244 and 1312 COVID-19 hospital admissions - dated between January 2020 and January 2021 from 10 health departments, were used respectively for training and validation of separate treatment-effect models (TE-ML) for remdesivir, corticosteroids, tocilizumab, lopinavir-ritonavir, azithromycin and chloroquine/hydroxychloroquine. 2390 admissions from 2 additional health departments were reserved as an independent test to analyze retrospectively the survival benefits of therapies in the population selected by the TE-ML models using cox-proportional hazard models. TE-ML models were adjusted using treatment propensity scores to control for pre-treatment confounding variables associated to outcome and further evaluated for futility. ML architecture was based on boosted decision-trees. Results: In the populations identified by the TE-ML models, only Remdesivir and Tocilizumab were significantly associated with an increase in survival time, with hazard ratios of 0.41 (P = 0.04) and 0.21 (P = 0.001), respectively. No survival benefits from chloroquine derivatives, lopinavir-ritonavir and azithromycin were demonstrated. Tools to explain the predictions of TE-ML models are explored at patient-level as potential tools for personalized decision making and precision medicine. Conclusion: ML methods are suitable tools toward RWE analysis of COVID-19 pharmacotherapies. Results obtained reproduce published results on RWE and validate the results from RCTs.
UMLS-ChestNet: A deep convolutional neural network for radiological findings, differential diagnoses and localizations of COVID-19 in chest x-rays
Jun 06, 2020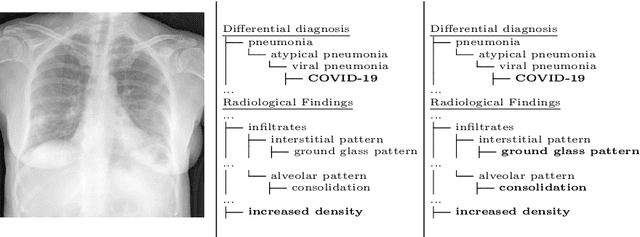
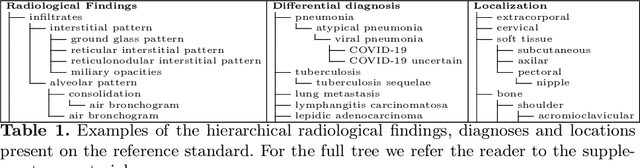


Abstract:In this work we present a method for the detection of radiological findings, their location and differential diagnoses from chest x-rays. Unlike prior works that focus on the detection of few pathologies, we use a hierarchical taxonomy mapped to the Unified Medical Language System (UMLS) terminology to identify 189 radiological findings, 22 differential diagnosis and 122 anatomic locations, including ground glass opacities, infiltrates, consolidations and other radiological findings compatible with COVID-19. We train the system on one large database of 92,594 frontal chest x-rays (AP or PA, standing, supine or decubitus) and a second database of 2,065 frontal images of COVID-19 patients identified by at least one positive Polymerase Chain Reaction (PCR) test. The reference labels are obtained through natural language processing of the radiological reports. On 23,159 test images, the proposed neural network obtains an AUC of 0.94 for the diagnosis of COVID-19. To our knowledge, this work uses the largest chest x-ray dataset of COVID-19 positive cases to date and is the first one to use a hierarchical labeling schema and to provide interpretability of the results, not only by using network attention methods, but also by indicating the radiological findings that have led to the diagnosis.
BIMCV COVID-19+: a large annotated dataset of RX and CT images from COVID-19 patients
Jun 05, 2020
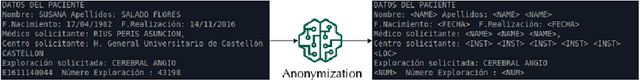

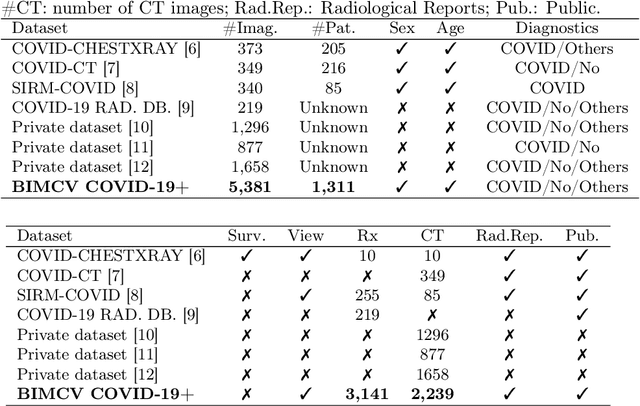

Abstract:This paper describes BIMCV COVID-19+, a large dataset from the Valencian Region Medical ImageBank (BIMCV) containing chest X-ray images CXR (CR, DX) and computed tomography (CT) imaging of COVID-19+ patients along with their radiological findings and locations, pathologies, radiological reports (in Spanish), DICOM metadata, Polymerase chain reaction (PCR), Immunoglobulin G (IgG) and Immunoglobulin M (IgM) diagnostic antibody tests. The findings have been mapped onto standard Unified Medical Language System (UMLS) terminology and cover a wide spectrum of thoracic entities, unlike the considerably more reduced number of entities annotated in previous datasets. Images are stored in high resolution and entities are localized with anatomical labels and stored in a Medical Imaging Data Structure (MIDS) format. In addition, 10 images were annotated by a team of radiologists to include semantic segmentation of radiological findings. This first iteration of the database includes 1,380 CX, 885 DX and 163 CT studies from 1,311 COVID-19+ patients. This is, to the best of our knowledge, the largest COVID-19+ dataset of images available in an open format. The dataset can be downloaded from http://bimcv.cipf.es/bimcv-projects/bimcv-covid19.
PadChest: A large chest x-ray image dataset with multi-label annotated reports
Feb 07, 2019

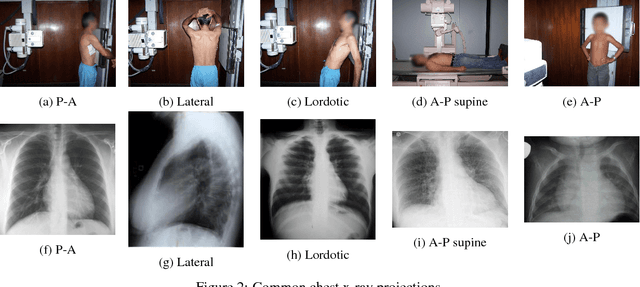
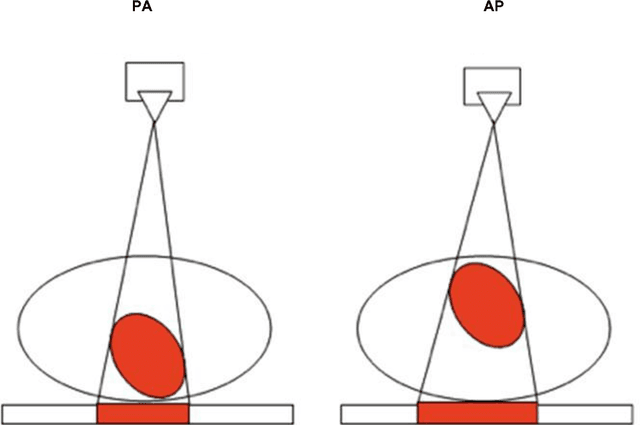
Abstract:We present a labeled large-scale, high resolution chest x-ray dataset for the automated exploration of medical images along with their associated reports. This dataset includes more than 160,000 images obtained from 67,000 patients that were interpreted and reported by radiologists at Hospital San Juan Hospital (Spain) from 2009 to 2017, covering six different position views and additional information on image acquisition and patient demography. The reports were labeled with 174 different radiographic findings, 19 differential diagnoses and 104 anatomic locations organized as a hierarchical taxonomy and mapped onto standard Unified Medical Language System (UMLS) terminology. Of these reports, 27% were manually annotated by trained physicians and the remaining set was labeled using a supervised method based on a recurrent neural network with attention mechanisms. The labels generated were then validated in an independent test set achieving a 0.93 Micro-F1 score. To the best of our knowledge, this is one of the largest public chest x-ray database suitable for training supervised models concerning radiographs, and the first to contain radiographic reports in Spanish. The PadChest dataset can be downloaded from http://bimcv.cipf.es/bimcv-projects/padchest/.
Learning Eligibility in Cancer Clinical Trials using Deep Neural Networks
Jul 25, 2018

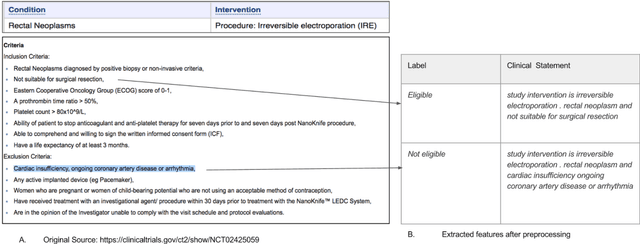

Abstract:Interventional cancer clinical trials are generally too restrictive, and some patients are often excluded on the basis of comorbidity, past or concomitant treatments, or the fact that they are over a certain age. The efficacy and safety of new treatments for patients with these characteristics are, therefore, not defined. In this work, we built a model to automatically predict whether short clinical statements were considered inclusion or exclusion criteria. We used protocols from cancer clinical trials that were available in public registries from the last 18 years to train word-embeddings, and we constructed a~dataset of 6M short free-texts labeled as eligible or not eligible. A text classifier was trained using deep neural networks, with pre-trained word-embeddings as inputs, to predict whether or not short free-text statements describing clinical information were considered eligible. We additionally analyzed the semantic reasoning of the word-embedding representations obtained and were able to identify equivalent treatments for a type of tumor analogous with the drugs used to treat other tumors. We show that representation learning using {deep} neural networks can be successfully leveraged to extract the medical knowledge from clinical trial protocols for potentially assisting practitioners when prescribing treatments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge