Anna Rieckmann
Detect and Correct Bias in Multi-Site Neuroimaging Datasets
Feb 12, 2020
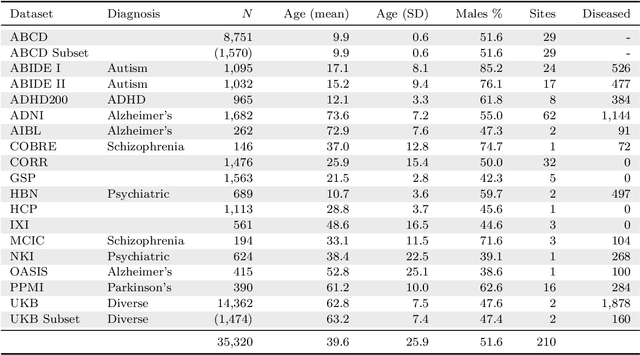
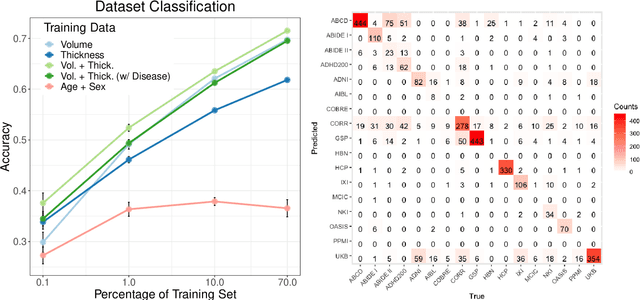
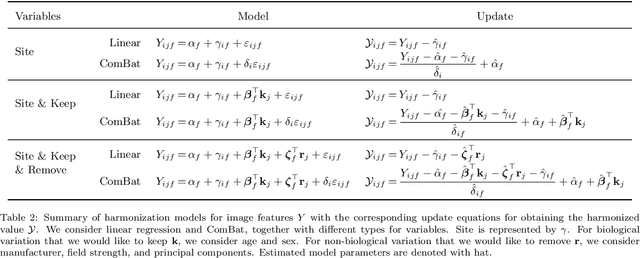
Abstract:The desire to train complex machine learning algorithms and to increase the statistical power in association studies drives neuroimaging research to use ever-larger datasets. The most obvious way to increase sample size is by pooling scans from independent studies. However, simple pooling is often ill-advised as selection, measurement, and confounding biases may creep in and yield spurious correlations. In this work, we combine 35,320 magnetic resonance images of the brain from 17 studies to examine bias in neuroimaging. In the first experiment, Name That Dataset, we provide empirical evidence for the presence of bias by showing that scans can be correctly assigned to their respective dataset with 71.5% accuracy. Given such evidence, we take a closer look at confounding bias, which is often viewed as the main shortcoming in observational studies. In practice, we neither know all potential confounders nor do we have data on them. Hence, we model confounders as unknown, latent variables. Kolmogorov complexity is then used to decide whether the confounded or the causal model provides the simplest factorization of the graphical model. Finally, we present methods for dataset harmonization and study their ability to remove bias in imaging features. In particular, we propose an extension of the recently introduced ComBat algorithm to control for global variation across image features, inspired by adjusting for population stratification in genetics. Our results demonstrate that harmonization can reduce dataset-specific information in image features. Further, confounding bias can be reduced and even turned into a causal relationship. However, harmonziation also requires caution as it can easily remove relevant subject-specific information.
Quantifying Confounding Bias in Neuroimaging Datasets with Causal Inference
Jul 09, 2019
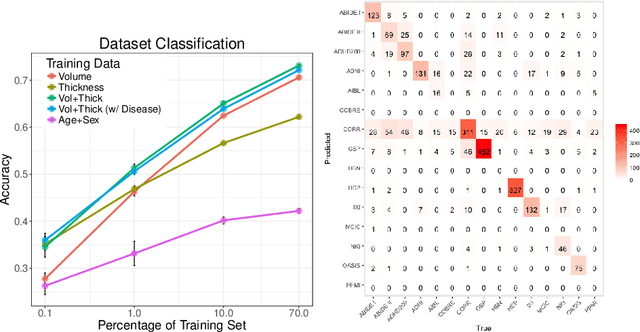
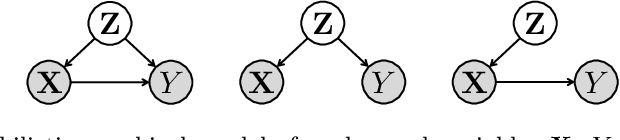
Abstract:Neuroimaging datasets keep growing in size to address increasingly complex medical questions. However, even the largest datasets today alone are too small for training complex machine learning models. A potential solution is to increase sample size by pooling scans from several datasets. In this work, we combine 12,207 MRI scans from 15 studies and show that simple pooling is often ill-advised due to introducing various types of biases in the training data. First, we systematically define these biases. Second, we detect bias by experimentally showing that scans can be correctly assigned to their respective dataset with 73.3% accuracy. Finally, we propose to tell causal from confounding factors by quantifying the extent of confounding and causality in a single dataset using causal inference. We achieve this by finding the simplest graphical model in terms of Kolmogorov complexity. As Kolmogorov complexity is not directly computable, we employ the minimum description length to approximate it. We empirically show that our approach is able to estimate plausible causal relationships from real neuroimaging data.
Detect, Quantify, and Incorporate Dataset Bias: A Neuroimaging Analysis on 12,207 Individuals
Apr 28, 2018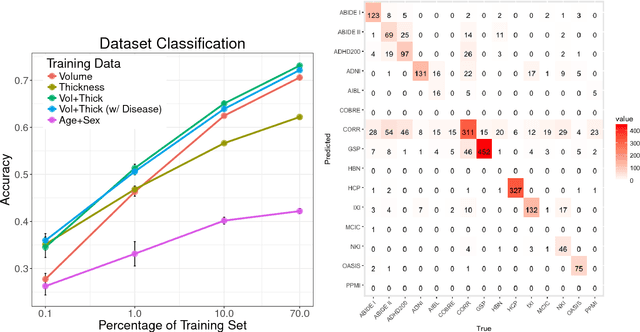
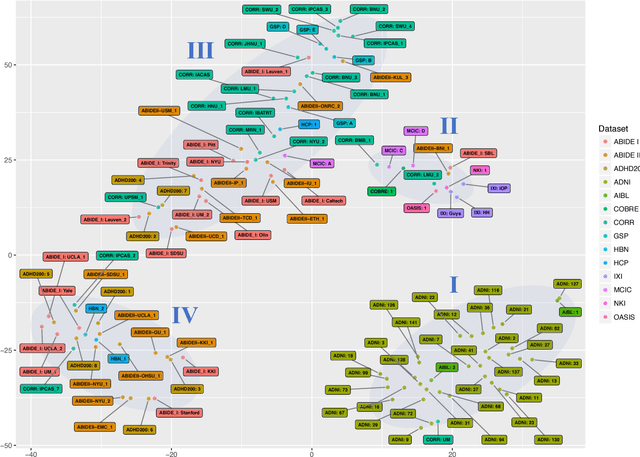
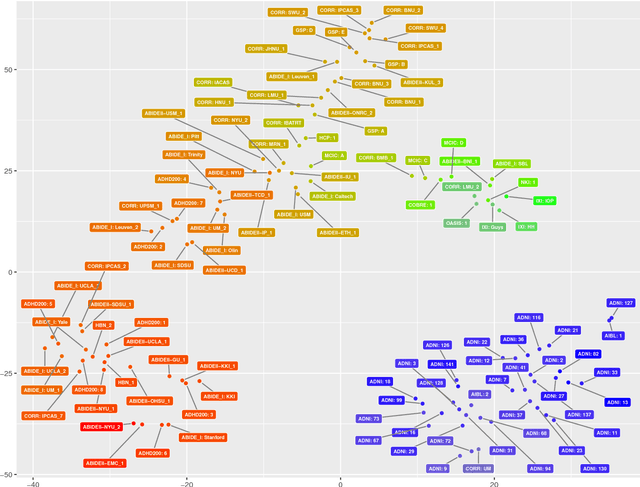
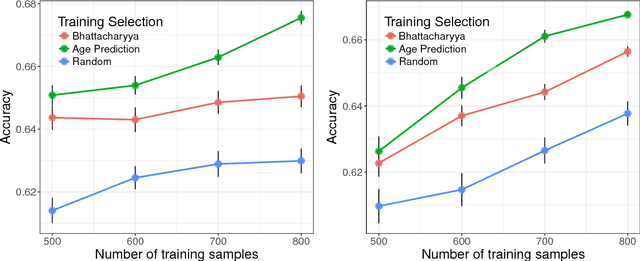
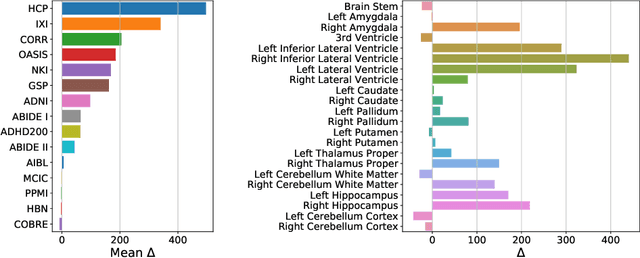
Abstract:Neuroimaging datasets keep growing in size to address increasingly complex medical questions. However, even the largest datasets today alone are too small for training complex models or for finding genome wide associations. A solution is to grow the sample size by merging data across several datasets. However, bias in datasets complicates this approach and includes additional sources of variation in the data instead. In this work, we combine 15 large neuroimaging datasets to study bias. First, we detect bias by demonstrating that scans can be correctly assigned to a dataset with 73.3% accuracy. Next, we introduce metrics to quantify the compatibility across datasets and to create embeddings of neuroimaging sites. Finally, we incorporate the presence of bias for the selection of a training set for predicting autism. For the quantification of the dataset bias, we introduce two metrics: the Bhattacharyya distance between datasets and the age prediction error. The presented embedding of neuroimaging sites provides an interesting new visualization about the similarity of different sites. This could be used to guide the merging of data sources, while limiting the introduction of unwanted variation. Finally, we demonstrate a clear performance increase when incorporating dataset bias for training set selection in autism prediction. Overall, we believe that the growing amount of neuroimaging data necessitates to incorporate data-driven methods for quantifying dataset bias in future analyses.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge