Andrew J. Reader
King's College London
Detecting and refurbishing ground truth errors during training of deep learning-based echocardiography segmentation models
Apr 14, 2026Abstract:Deep learning-based medical image segmentation typically relies on ground truth (GT) labels obtained through manual annotation, but these can be prone to random errors or systematic biases. This study examines the robustness of deep learning models to such errors in echocardiography (echo) segmentation and evaluates a novel strategy for detecting and refurbishing erroneous labels during model training. Using the CAMUS dataset, we simulate three error types, then compare a loss-based GT label error detection method with one based on Variance of Gradients (VOG). We also propose a pseudo-labelling approach to refurbish suspected erroneous GT labels. We assess the performance of our proposed approach under varying error levels. Results show that VOG proved highly effective in flagging erroneous GT labels during training. However, a standard U-Net maintained strong performance under random label errors and moderate levels of systematic errors (up to 50%). The detection and refurbishment approach improved performance, particularly under high-error conditions.
Personalized MR-Informed Diffusion Models for 3D PET Image Reconstruction
Jun 04, 2025Abstract:Recent work has shown improved lesion detectability and flexibility to reconstruction hyperparameters (e.g. scanner geometry or dose level) when PET images are reconstructed by leveraging pre-trained diffusion models. Such methods train a diffusion model (without sinogram data) on high-quality, but still noisy, PET images. In this work, we propose a simple method for generating subject-specific PET images from a dataset of multi-subject PET-MR scans, synthesizing "pseudo-PET" images by transforming between different patients' anatomy using image registration. The images we synthesize retain information from the subject's MR scan, leading to higher resolution and the retention of anatomical features compared to the original set of PET images. With simulated and real [$^{18}$F]FDG datasets, we show that pre-training a personalized diffusion model with subject-specific "pseudo-PET" images improves reconstruction accuracy with low-count data. In particular, the method shows promise in combining information from a guidance MR scan without overly imposing anatomical features, demonstrating an improved trade-off between reconstructing PET-unique image features versus features present in both PET and MR. We believe this approach for generating and utilizing synthetic data has further applications to medical imaging tasks, particularly because patient-specific PET images can be generated without resorting to generative deep learning or large training datasets.
Multi-Subject Image Synthesis as a Generative Prior for Single-Subject PET Image Reconstruction
Dec 05, 2024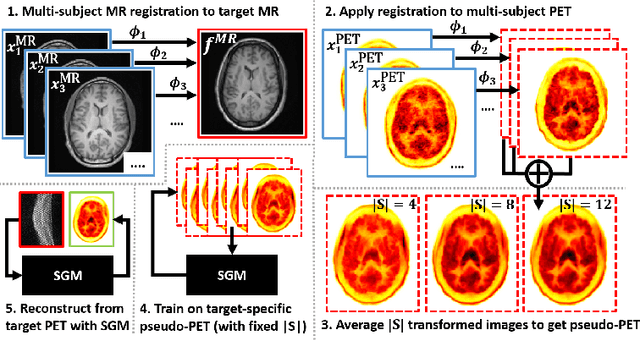
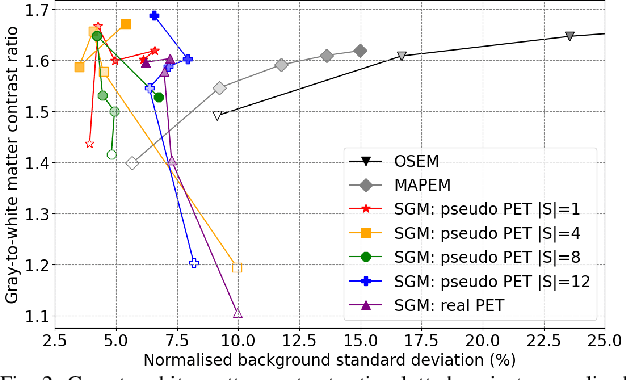
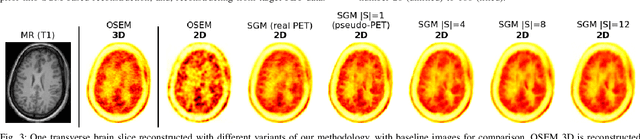
Abstract:Large high-quality medical image datasets are difficult to acquire but necessary for many deep learning applications. For positron emission tomography (PET), reconstructed image quality is limited by inherent Poisson noise. We propose a novel method for synthesising diverse and realistic pseudo-PET images with improved signal-to-noise ratio. We also show how our pseudo-PET images may be exploited as a generative prior for single-subject PET image reconstruction. Firstly, we perform deep-learned deformable registration of multi-subject magnetic resonance (MR) images paired to multi-subject PET images. We then use the anatomically-learned deformation fields to transform multiple PET images to the same reference space, before averaging random subsets of the transformed multi-subject data to form a large number of varying pseudo-PET images. We observe that using MR information for registration imbues the resulting pseudo-PET images with improved anatomical detail compared to the originals. We consider applications to PET image reconstruction, by generating pseudo-PET images in the same space as the intended single-subject reconstruction and using them as training data for a diffusion model-based reconstruction method. We show visual improvement and reduced background noise in our 2D reconstructions as compared to OSEM, MAP-EM and an existing state-of-the-art diffusion model-based approach. Our method shows the potential for utilising highly subject-specific prior information within a generative reconstruction framework. Future work may compare the benefits of our approach to explicitly MR-guided reconstruction methodologies.
Likelihood-Scheduled Score-Based Generative Modeling for Fully 3D PET Image Reconstruction
Dec 05, 2024Abstract:Medical image reconstruction with pre-trained score-based generative models (SGMs) has advantages over other existing state-of-the-art deep-learned reconstruction methods, including improved resilience to different scanner setups and advanced image distribution modeling. SGM-based reconstruction has recently been applied to simulated positron emission tomography (PET) datasets, showing improved contrast recovery for out-of-distribution lesions relative to the state-of-the-art. However, existing methods for SGM-based reconstruction from PET data suffer from slow reconstruction, burdensome hyperparameter tuning and slice inconsistency effects (in 3D). In this work, we propose a practical methodology for fully 3D reconstruction that accelerates reconstruction and reduces the number of critical hyperparameters by matching the likelihood of an SGM's reverse diffusion process to a current iterate of the maximum-likelihood expectation maximization algorithm. Using the example of low-count reconstruction from simulated $[^{18}$F]DPA-714 datasets, we show our methodology can match or improve on the NRMSE and SSIM of existing state-of-the-art SGM-based PET reconstruction while reducing reconstruction time and the need for hyperparameter tuning. We evaluate our methodology against state-of-the-art supervised and conventional reconstruction algorithms. Finally, we demonstrate a first-ever implementation of SGM-based reconstruction for real 3D PET data, specifically $[^{18}$F]DPA-714 data, where we integrate perpendicular pre-trained SGMs to eliminate slice inconsistency issues.
Generative-Model-Based Fully 3D PET Image Reconstruction by Conditional Diffusion Sampling
Dec 05, 2024Abstract:Score-based generative models (SGMs) have recently shown promising results for image reconstruction on simulated positron emission tomography (PET) datasets. In this work we have developed and implemented practical methodology for 3D image reconstruction with SGMs, and perform (to our knowledge) the first SGM-based reconstruction of real fully 3D PET data. We train an SGM on full-count reference brain images, and extend methodology to allow SGM-based reconstructions at very low counts (1% of original, to simulate low-dose or short-duration scanning). We then perform reconstructions for multiple independent realisations of 1% count data, allowing us to analyse the bias and variance characteristics of the method. We sample from the learned posterior distribution of the generative algorithm to calculate uncertainty images for our reconstructions. We evaluate the method's performance on real full- and low-count PET data and compare with conventional OSEM and MAP-EM baselines, showing that our SGM-based low-count reconstructions match full-dose reconstructions more closely and in a bias-variance trade-off comparison, our SGM-reconstructed images have lower variance than existing baselines. Future work will compare to supervised deep-learned methods, with other avenues for investigation including how data conditioning affects the SGM's posterior distribution and the algorithm's performance with different tracers.
Label Dropout: Improved Deep Learning Echocardiography Segmentation Using Multiple Datasets With Domain Shift and Partial Labelling
Mar 12, 2024Abstract:Echocardiography (echo) is the first imaging modality used when assessing cardiac function. The measurement of functional biomarkers from echo relies upon the segmentation of cardiac structures and deep learning models have been proposed to automate the segmentation process. However, in order to translate these tools to widespread clinical use it is important that the segmentation models are robust to a wide variety of images (e.g. acquired from different scanners, by operators with different levels of expertise etc.). To achieve this level of robustness it is necessary that the models are trained with multiple diverse datasets. A significant challenge faced when training with multiple diverse datasets is the variation in label presence, i.e. the combined data are often partially-labelled. Adaptations of the cross entropy loss function have been proposed to deal with partially labelled data. In this paper we show that training naively with such a loss function and multiple diverse datasets can lead to a form of shortcut learning, where the model associates label presence with domain characteristics, leading to a drop in performance. To address this problem, we propose a novel label dropout scheme to break the link between domain characteristics and the presence or absence of labels. We demonstrate that label dropout improves echo segmentation Dice score by 62% and 25% on two cardiac structures when training using multiple diverse partially labelled datasets.
Self-Supervised and Supervised Deep Learning for PET Image Reconstruction
Feb 25, 2023



Abstract:A unified self-supervised and supervised deep learning framework for PET image reconstruction is presented, including deep-learned filtered backprojection (DL-FBP) for sinograms, deep-learned backproject then filter (DL-BPF) for backprojected images, and a more general mapping using a deep network in both the sinogram and image domains (DL-FBP-F). The framework allows varying amounts and types of training data, from the case of having only one single dataset to reconstruct through to the case of having numerous measured datasets, which may or may not be paired with high-quality references. For purely self-supervised mappings, no reference or ground truth data are needed. The self-supervised deep-learned reconstruction operators all use a conventional image reconstruction objective within the loss function (e.g. maximum Poisson likelihood, maximum a posteriori). If it is desired for the reconstruction networks to generalise (i.e. to need either no or minimal retraining for a new measured dataset, but to be fast, ready to reuse), then these self-supervised networks show potential even when previously trained from just one single dataset. For any given new measured dataset, finetuning is however often necessary, and of course the initial training set should ideally go beyond just one dataset if a generalisable network is sought. Example results for the purely self-supervised single-dataset case are shown, but the networks can be i) trained uniquely for any measured dataset to reconstruct from, ii) pretrained on multiple datasets and then used with no retraining for new measured data, iii) pretrained and then finetuned for new measured data, iv) optionally trained with high-quality references. The framework, with its optional inclusion of supervised learning, provides a spectrum of reconstruction approaches by making use of whatever (if any) training data quantities and types are available.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge