Alaa Abi-Haidar
Collective Classification of Textual Documents by Guided Self-Organization in T-Cell Cross-Regulation Dynamics
Feb 04, 2011
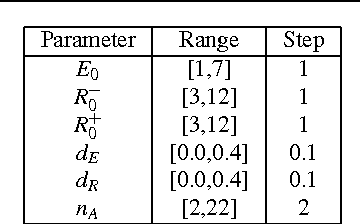


Abstract:We present and study an agent-based model of T-Cell cross-regulation in the adaptive immune system, which we apply to binary classification. Our method expands an existing analytical model of T-cell cross-regulation (Carneiro et al. in Immunol Rev 216(1):48-68, 2007) that was used to study the self-organizing dynamics of a single population of T-Cells in interaction with an idealized antigen presenting cell capable of presenting a single antigen. With agent-based modeling we are able to study the self-organizing dynamics of multiple populations of distinct T-cells which interact via antigen presenting cells that present hundreds of distinct antigens. Moreover, we show that such self-organizing dynamics can be guided to produce an effective binary classification of antigens, which is competitive with existing machine learning methods when applied to biomedical text classification. More specifically, here we test our model on a dataset of publicly available full-text biomedical articles provided by the BioCreative challenge (Krallinger in The biocreative ii. 5 challenge overview, p 19, 2009). We study the robustness of our model's parameter configurations, and show that it leads to encouraging results comparable to state-of-the-art classifiers. Our results help us understand both T-cell cross-regulation as a general principle of guided self-organization, as well as its applicability to document classification. Therefore, we show that our bio-inspired algorithm is a promising novel method for biomedical article classification and for binary document classification in general.
Uncovering protein interaction in abstracts and text using a novel linear model and word proximity networks
Dec 04, 2008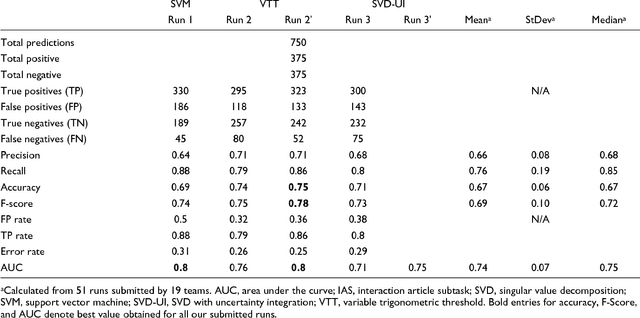
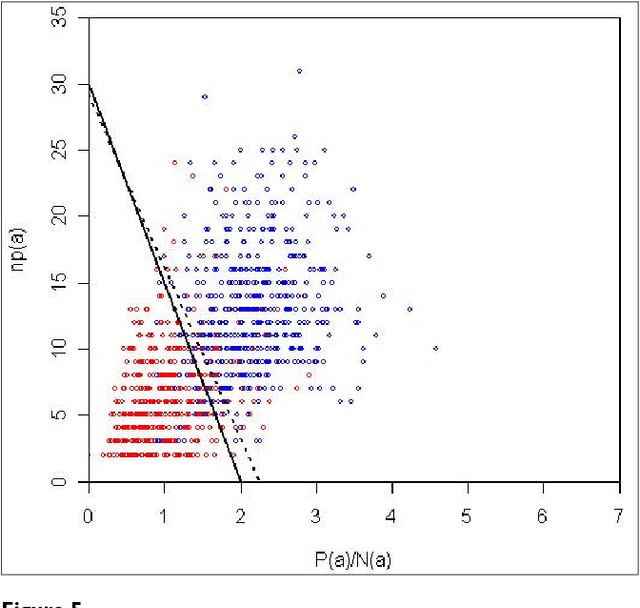
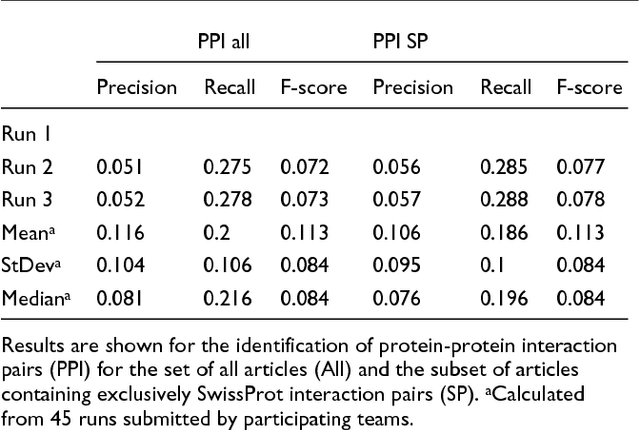
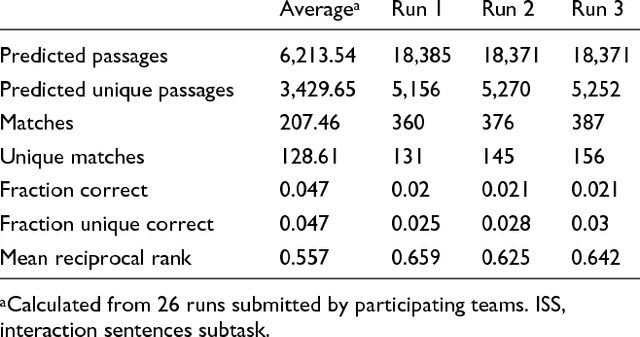
Abstract:We participated in three of the protein-protein interaction subtasks of the Second BioCreative Challenge: classification of abstracts relevant for protein-protein interaction (IAS), discovery of protein pairs (IPS) and text passages characterizing protein interaction (ISS) in full text documents. We approached the abstract classification task with a novel, lightweight linear model inspired by spam-detection techniques, as well as an uncertainty-based integration scheme. We also used a Support Vector Machine and the Singular Value Decomposition on the same features for comparison purposes. Our approach to the full text subtasks (protein pair and passage identification) includes a feature expansion method based on word-proximity networks. Our approach to the abstract classification task (IAS) was among the top submissions for this task in terms of the measures of performance used in the challenge evaluation (accuracy, F-score and AUC). We also report on a web-tool we produced using our approach: the Protein Interaction Abstract Relevance Evaluator (PIARE). Our approach to the full text tasks resulted in one of the highest recall rates as well as mean reciprocal rank of correct passages. Our approach to abstract classification shows that a simple linear model, using relatively few features, is capable of generalizing and uncovering the conceptual nature of protein-protein interaction from the bibliome. Since the novel approach is based on a very lightweight linear model, it can be easily ported and applied to similar problems. In full text problems, the expansion of word features with word-proximity networks is shown to be useful, though the need for some improvements is discussed.
Adaptive Spam Detection Inspired by a Cross-Regulation Model of Immune Dynamics: A Study of Concept Drift
Dec 04, 2008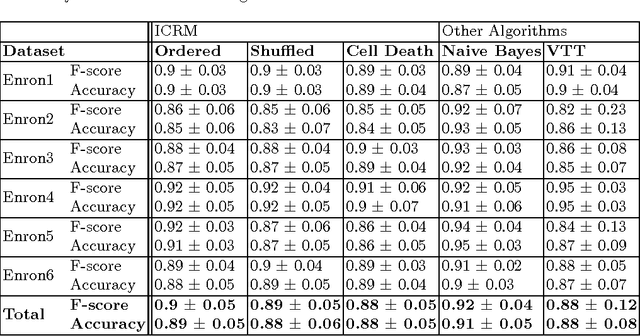



Abstract:This paper proposes a novel solution to spam detection inspired by a model of the adaptive immune system known as the crossregulation model. We report on the testing of a preliminary algorithm on six e-mail corpora. We also compare our results statically and dynamically with those obtained by the Naive Bayes classifier and another binary classification method we developed previously for biomedical text-mining applications. We show that the cross-regulation model is competitive against those and thus promising as a bio-inspired algorithm for spam detection in particular, and binary classification in general.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge