TF3P: Three-dimensional Force Fields Fingerprint Learned by Deep Capsular Network
Paper and Code
Dec 25, 2019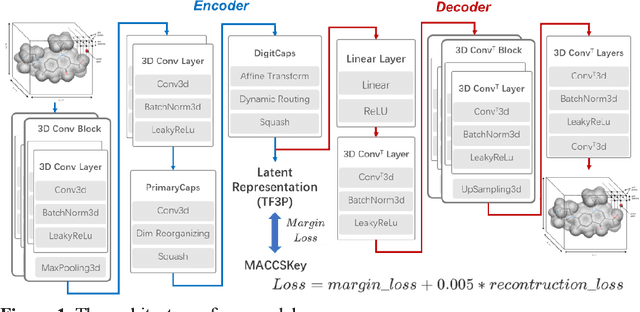


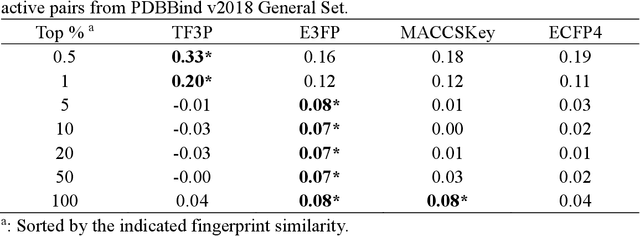
Molecular fingerprints are the workhorse in ligand-based drug discovery. In recent years, increasing number of research papers reported fascinating results on using deep neural networks to learn 2D molecular representations as fingerprints. One may anticipate that the integration of deep learning would also contribute to the prosperity of 3D fingerprints. Here, we presented a new 3D small molecule fingerprint, the three-dimensional force fields fingerprint (TF3P), learned by deep capsular network whose training is in no need of labeled dataset for specific predictive tasks. TF3P can encode the 3D force fields information of molecules and demonstrates its stronger ability to capture 3D structural changes, recognize molecules alike in 3D but not in 2D, and recognize similar targets inaccessible by other fingerprints, including the solely existing 3D fingerprint E3FP, based on only ligands similarity. Furthermore, TF3P is compatible with both statistical models (e.g. similarity ensemble approach) and machine learning models. Altogether, we report TF3P as a new 3D small molecule fingerprint with promising future in ligand-based drug discovery.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge