Spiral Contrastive Learning: An Efficient 3D Representation Learning Method for Unannotated CT Lesions
Paper and Code
Aug 23, 2022
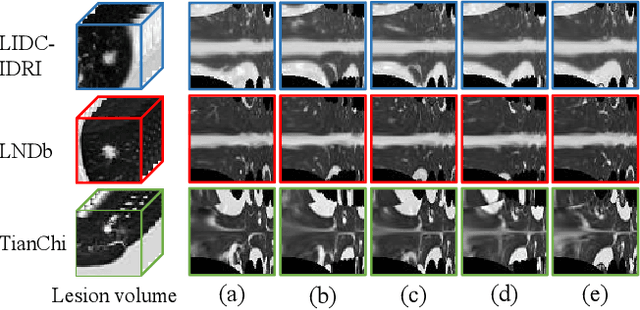
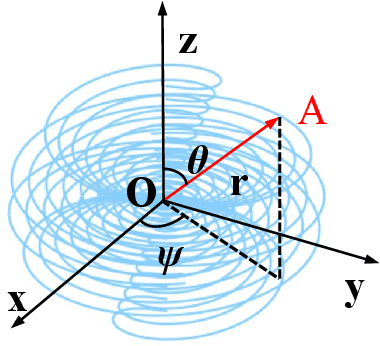

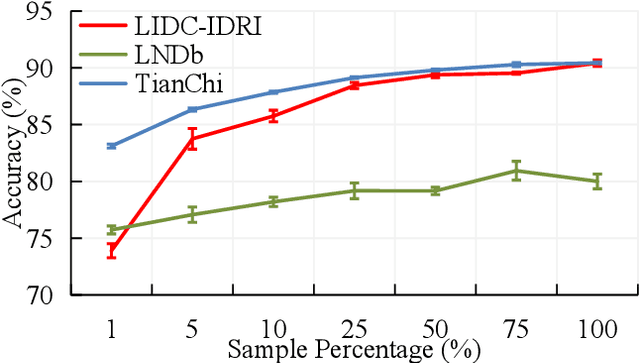
Computed tomography (CT) samples with pathological annotations are difficult to obtain. As a result, the computer-aided diagnosis (CAD) algorithms are trained on small datasets (e.g., LIDC-IDRI with 1,018 samples), limiting their accuracies and reliability. In the past five years, several works have tailored for unsupervised representations of CT lesions via two-dimensional (2D) and three-dimensional (3D) self-supervised learning (SSL) algorithms. The 2D algorithms have difficulty capturing 3D information, and existing 3D algorithms are computationally heavy. Light-weight 3D SSL remains the boundary to explore. In this paper, we propose the spiral contrastive learning (SCL), which yields 3D representations in a computationally efficient manner. SCL first transforms 3D lesions to the 2D plane using an information-preserving spiral transformation, and then learn transformation-invariant features using 2D contrastive learning. For the augmentation, we consider natural image augmentations and medical image augmentations. We evaluate SCL by training a classification head upon the embedding layer. Experimental results show that SCL achieves state-of-the-art accuracy on LIDC-IDRI (89.72%), LNDb (82.09%) and TianChi (90.16%) for unsupervised representation learning. With 10% annotated data for fine-tune, the performance of SCL is comparable to that of supervised learning algorithms (85.75% vs. 85.03% on LIDC-IDRI, 78.20% vs. 73.44% on LNDb and 87.85% vs. 83.34% on TianChi, respectively). Meanwhile, SCL reduces the computational effort by 66.98% compared to other 3D SSL algorithms, demonstrating the efficiency of the proposed method in unsupervised pre-training.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge