Self-Rule to Adapt: Generalized Multi-source Feature Learning Using Unsupervised Domain Adaptation for Colorectal Cancer Tissue Detection
Paper and Code
Aug 20, 2021
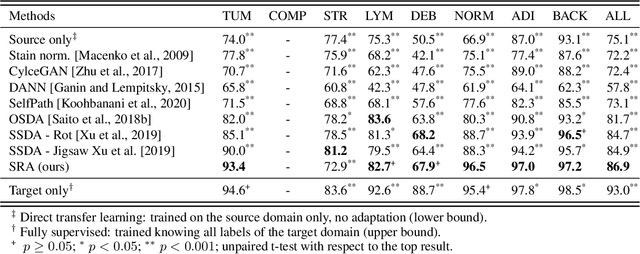
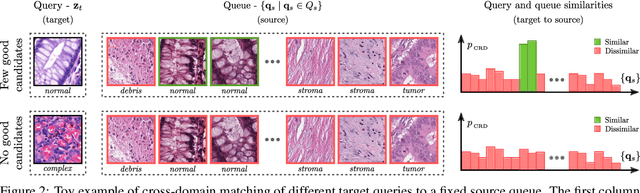
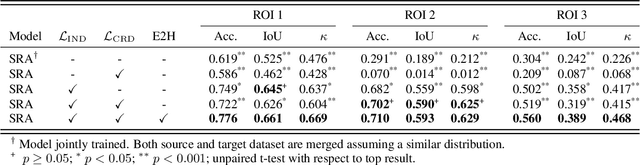
Supervised learning is constrained by the availability of labeled data, which are especially expensive to acquire in the field of digital pathology. Making use of open-source data for pre-training or using domain adaptation can be a way to overcome this issue. However, pre-trained networks often fail to generalize to new test domains that are not distributed identically due to variations in tissue stainings, types, and textures. Additionally, current domain adaptation methods mainly rely on fully-labeled source datasets. In this work, we propose SRA, which takes advantage of self-supervised learning to perform domain adaptation and removes the necessity of a fully-labeled source dataset. SRA can effectively transfer the discriminative knowledge obtained from a few labeled source domain's data to a new target domain without requiring additional tissue annotations. Our method harnesses both domains' structures by capturing visual similarity with intra-domain and cross-domain self-supervision. Moreover, we present a generalized formulation of our approach that allows the architecture to learn from multi-source domains. We show that our proposed method outperforms baselines for domain adaptation of colorectal tissue type classification and further validate our approach on our in-house clinical cohort. The code and models are available open-source: https://github.com/christianabbet/SRA.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge