PIVETed-Granite: Computational Phenotypes through Constrained Tensor Factorization
Paper and Code
Aug 08, 2018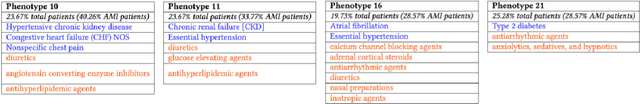

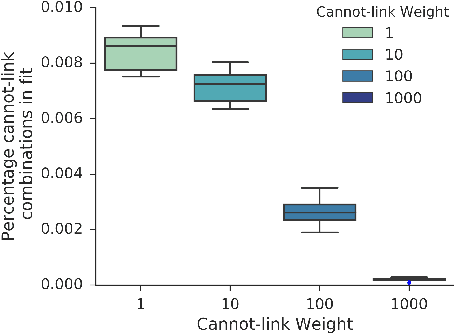

It has been recently shown that sparse, nonnegative tensor factorization of multi-modal electronic health record data is a promising approach to high-throughput computational phenotyping. However, such approaches typically do not leverage available domain knowledge while extracting the phenotypes; hence, some of the suggested phenotypes may not map well to clinical concepts or may be very similar to other suggested phenotypes. To address these issues, we present a novel, automatic approach called PIVETed-Granite that mines existing biomedical literature (PubMed) to obtain cannot-link constraints that are then used as side-information during a tensor-factorization based computational phenotyping process. The resulting improvements are clearly observed in experiments using a large dataset from VUMC to identify phenotypes for hypertensive patients.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge