Node-Aligned Graph-to-Graph Generation for Retrosynthesis Prediction
Paper and Code
Sep 27, 2023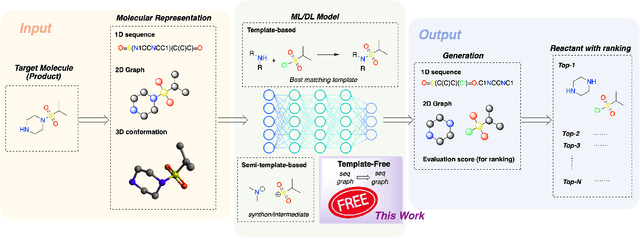

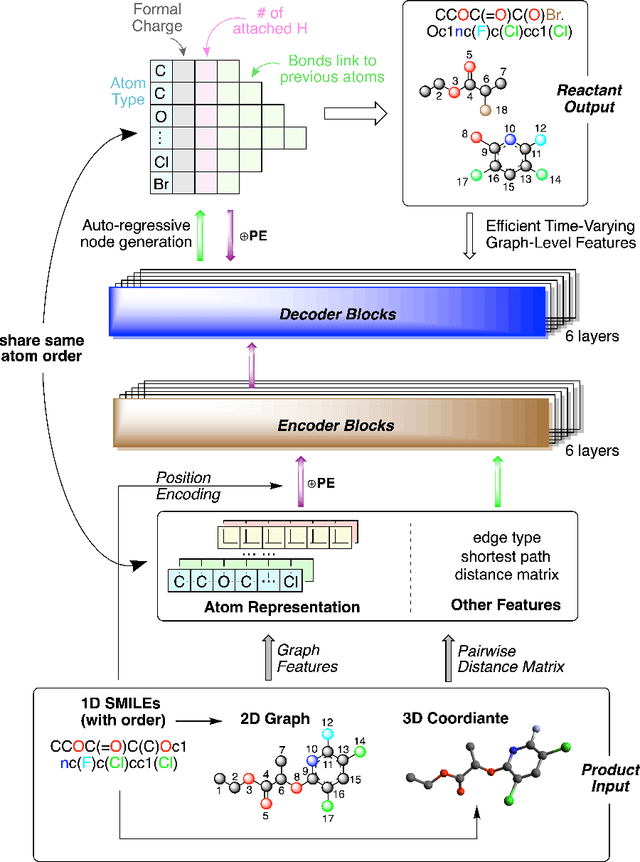

Single-step retrosynthesis is a crucial task in organic chemistry and drug design, requiring the identification of required reactants to synthesize a specific compound. with the advent of computer-aided synthesis planning, there is growing interest in using machine-learning techniques to facilitate the process. Existing template-free machine learning-based models typically utilize transformer structures and represent molecules as ID sequences. However, these methods often face challenges in fully leveraging the extensive topological information of the molecule and aligning atoms between the production and reactants, leading to results that are not as competitive as those of semi-template models. Our proposed method, Node-Aligned Graph-to-Graph (NAG2G), also serves as a transformer-based template-free model but utilizes 2D molecular graphs and 3D conformation information. Furthermore, our approach simplifies the incorporation of production-reactant atom mapping alignment by leveraging node alignment to determine a specific order for node generation and generating molecular graphs in an auto-regressive manner node-by-node. This method ensures that the node generation order coincides with the node order in the input graph, overcoming the difficulty of determining a specific node generation order in an auto-regressive manner. Our extensive benchmarking results demonstrate that the proposed NAG2G can outperform the previous state-of-the-art baselines in various metrics.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge