MedDistant19: A Challenging Benchmark for Distantly Supervised Biomedical Relation Extraction
Paper and Code
Apr 10, 2022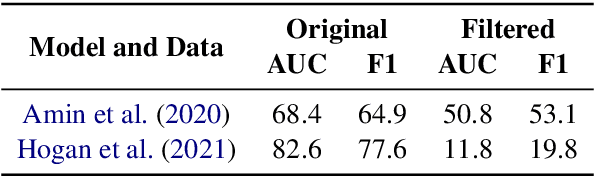
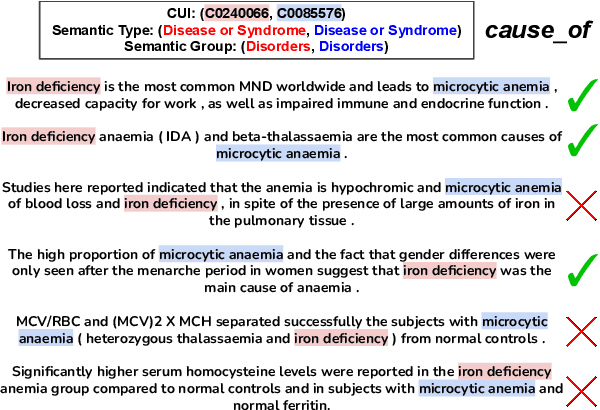

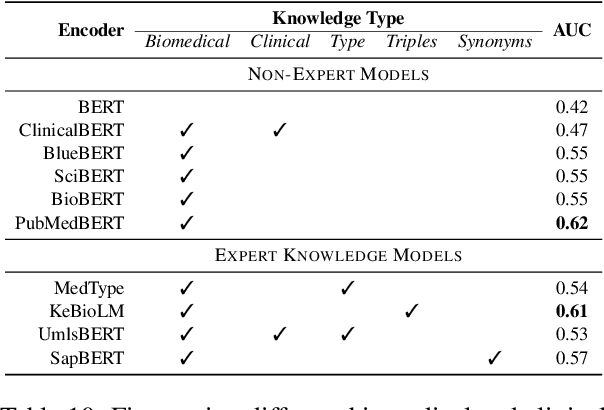

Relation Extraction in the biomedical domain is challenging due to the lack of labeled data and high annotation costs, needing domain experts. Distant supervision is commonly used as a way to tackle the scarcity of annotated data by automatically pairing knowledge graph relationships with raw texts. Distantly Supervised Biomedical Relation Extraction (Bio-DSRE) models can seemingly produce very accurate results in several benchmarks. However, given the challenging nature of the task, we set out to investigate the validity of such impressive results. We probed the datasets used by Amin et al. (2020) and Hogan et al. (2021) and found a significant overlap between training and evaluation relationships that, once resolved, reduced the accuracy of the models by up to 71%. Furthermore, we noticed several inconsistencies with the data construction process, such as creating negative samples and improper handling of redundant relationships. We mitigate these issues and present MedDistant19, a new benchmark dataset obtained by aligning the MEDLINE abstracts with the widely used SNOMED Clinical Terms (SNOMED CT) knowledge base. We experimented with several state-of-the-art models achieving an AUC of 55.4% and 49.8% at sentence- and bag-level, showing that there is still plenty of room for improvement.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge