MDM: Molecular Diffusion Model for 3D Molecule Generation
Paper and Code
Sep 13, 2022
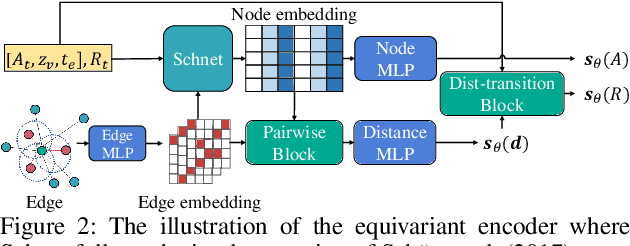
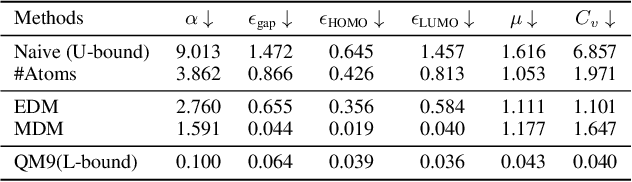
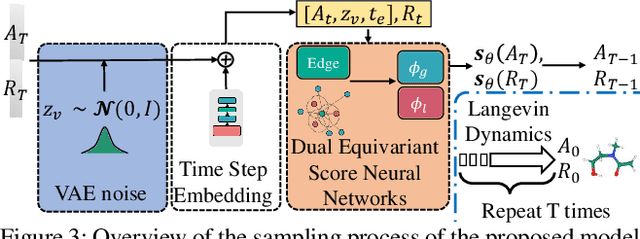
Molecule generation, especially generating 3D molecular geometries from scratch (i.e., 3D \textit{de novo} generation), has become a fundamental task in drug designs. Existing diffusion-based 3D molecule generation methods could suffer from unsatisfactory performances, especially when generating large molecules. At the same time, the generated molecules lack enough diversity. This paper proposes a novel diffusion model to address those two challenges. First, interatomic relations are not in molecules' 3D point cloud representations. Thus, it is difficult for existing generative models to capture the potential interatomic forces and abundant local constraints. To tackle this challenge, we propose to augment the potential interatomic forces and further involve dual equivariant encoders to encode interatomic forces of different strengths. Second, existing diffusion-based models essentially shift elements in geometry along the gradient of data density. Such a process lacks enough exploration in the intermediate steps of the Langevin dynamics. To address this issue, we introduce a distributional controlling variable in each diffusion/reverse step to enforce thorough explorations and further improve generation diversity. Extensive experiments on multiple benchmarks demonstrate that the proposed model significantly outperforms existing methods for both unconditional and conditional generation tasks. We also conduct case studies to help understand the physicochemical properties of the generated molecules.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge