Learning Rotation Invariant Features for Cryogenic Electron Microscopy Image Reconstruction
Paper and Code
Jan 10, 2021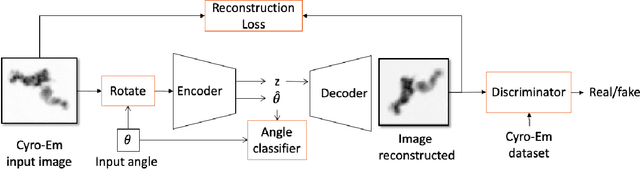
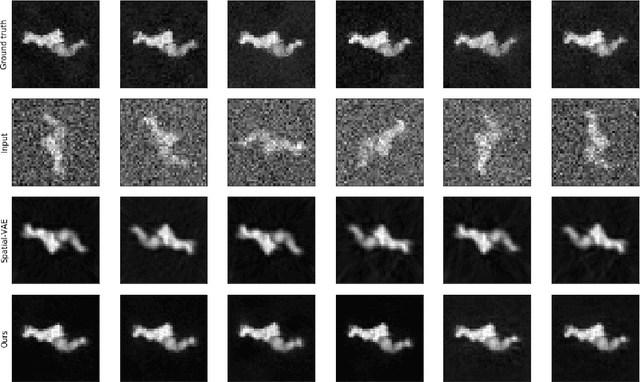
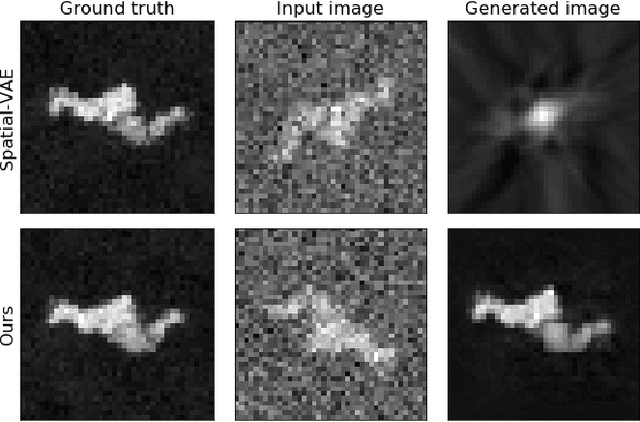
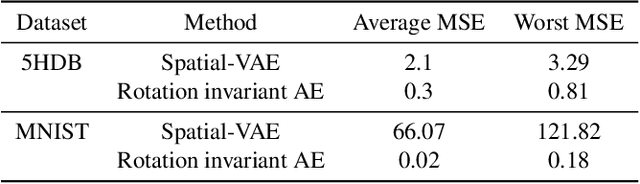


Cryo-Electron Microscopy (Cryo-EM) is a Nobel prize-winning technology for determining the 3D structure of particles at near-atomic resolution. A fundamental step in the recovering of the 3D single-particle structure is to align its 2D projections; thus, the construction of a canonical representation with a fixed rotation angle is required. Most approaches use discrete clustering which fails to capture the continuous nature of image rotation, others suffer from low-quality image reconstruction. We propose a novel method that leverages the recent development in the generative adversarial networks. We introduce an encoder-decoder with a rotation angle classifier. In addition, we utilize a discriminator on the decoder output to minimize the reconstruction error. We demonstrate our approach with the Cryo-EM 5HDB and the rotated MNIST datasets showing substantial improvement over recent methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge