Learning New Tricks from Old Dogs -- Inter-Species, Inter-Tissue Domain Adaptation for Mitotic Figure Assessment
Paper and Code
Nov 25, 2019
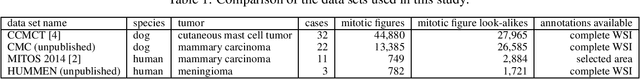
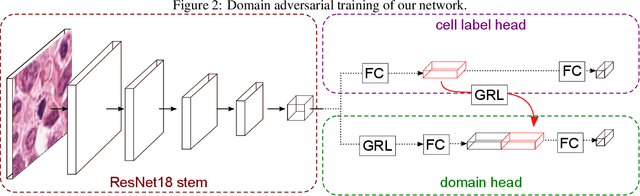
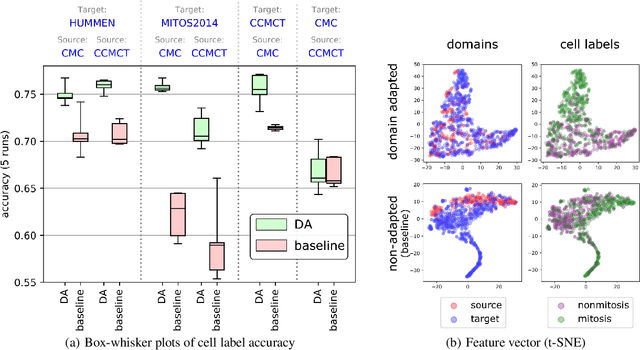
For histopathological tumor assessment, the count of mitotic figures per area is an important part of prognostication. Algorithmic approaches - such as for mitotic figure identification - have significantly improved in recent times, potentially allowing for computer-augmented or fully automatic screening systems in the future. This trend is further supported by whole slide scanning microscopes becoming available in many pathology labs and could soon become a standard imaging tool. For an application in broader fields of such algorithms, the availability of mitotic figure data sets of sufficient size for the respective tissue type and species is an important precondition, that is, however, rarely met. While algorithmic performance climbed steadily for e.g. human mammary carcinoma, thanks to several challenges held in the field, for most tumor types, data sets are not available. In this work, we assess domain transfer of mitotic figure recognition using domain adversarial training on four data sets, two from dogs and two from humans. We were able to show that domain adversarial training considerably improves accuracy when applying mitotic figure classification learned from the canine on the human data sets (up to +12.8% in accuracy) and is thus a helpful method to transfer knowledge from existing data sets to new tissue types and species.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge