Hierarchical Graph Convolutional Network Built by Multiscale Atlases for Brain Disorder Diagnosis Using Functional Connectivity
Paper and Code
Sep 22, 2022

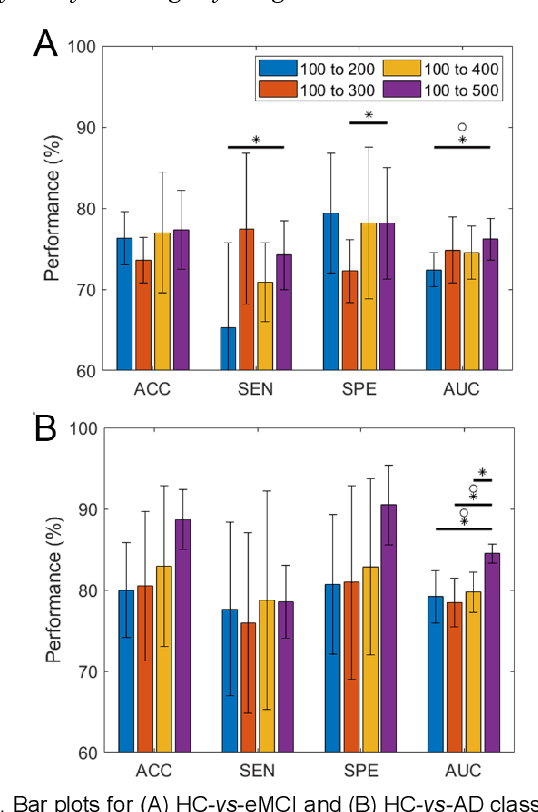
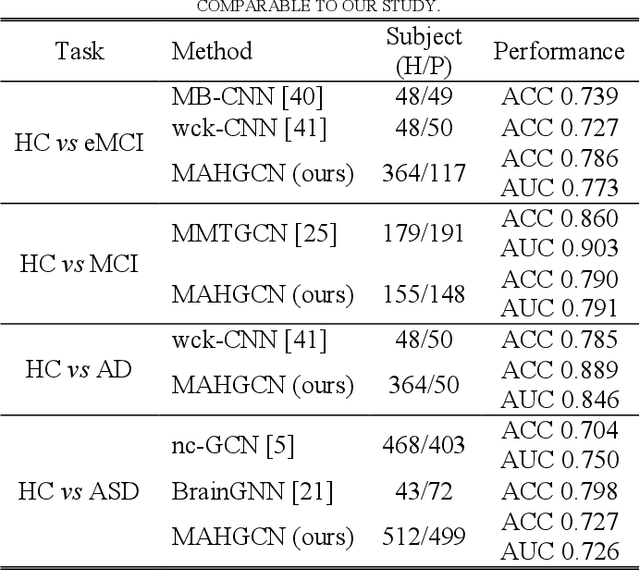
Functional connectivity network (FCN) data from functional magnetic resonance imaging (fMRI) is increasingly used for the diagnoses of brain disorders. However, state-of-the-art studies used to build the FCN using a single brain parcellation atlas at a certain spatial scale, which largely neglected functional interactions across different spatial scales in hierarchical manners. In this study, we propose a novel framework to perform multiscale FCN analysis for brain disorder diagnosis. We first use a set of well-defined multiscale atlases to compute multiscale FCNs. Then, we utilize biologically meaningful brain hierarchical relationships among the regions in multiscale atlases to perform nodal pooling across multiple spatial scales, namely "Atlas-guided Pooling". Accordingly, we propose a Multiscale-Atlases-based Hierarchical Graph Convolutional Network (MAHGCN), built on the stacked layers of graph convolution and the atlas-guided pooling, for a comprehensive extraction of diagnostic information from multiscale FCNs. Experiments on neuroimaging data from 1792 subjects demonstrate the effectiveness of our proposed method in the diagnoses of Alzheimer's disease (AD), the prodromal stage of AD (i.e., mild cognitive impairment [MCI]), as well as autism spectrum disorder (ASD), with accuracy of 88.9%, 78.6%, and 72.7% respectively. All results show significant advantages of our proposed method over other competing methods. This study not only demonstrates the feasibility of brain disorder diagnosis using resting-state fMRI empowered by deep learning, but also highlights that the functional interactions in the multiscale brain hierarchy are worth being explored and integrated into deep learning network architectures for better understanding the neuropathology of brain disorders.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge