Deep Feature based Cross-slide Registration
Paper and Code
Feb 27, 2022
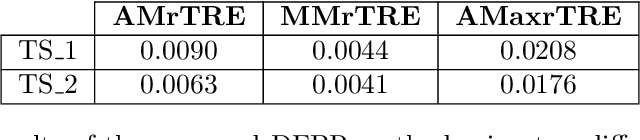
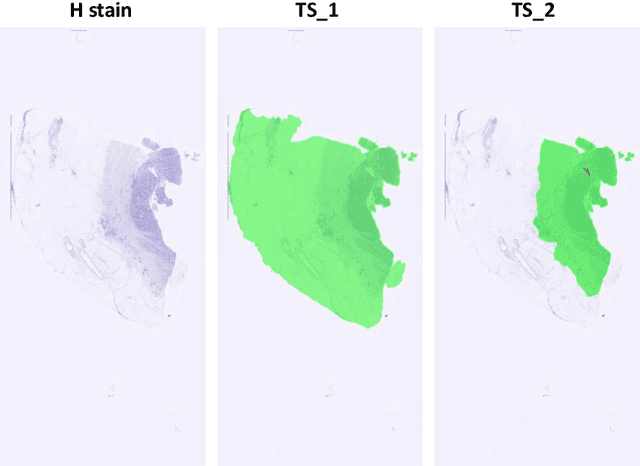
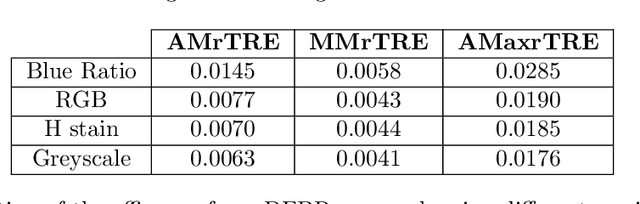
Cross-slide image analysis provides additional information by analysing the expression of different biomarkers as compared to a single slide analysis. Slides stained with different biomarkers are analysed side by side which may reveal unknown relations between the different biomarkers. During the slide preparation, a tissue section may be placed at an arbitrary orientation as compared to other sections of the same tissue block. The problem is compounded by the fact that tissue contents are likely to change from one section to the next and there may be unique artefacts on some of the slides. This makes registration of each section to a reference section of the same tissue block an important pre-requisite task before any cross-slide analysis. We propose a deep feature based registration (DFBR) method which utilises data-driven features to estimate the rigid transformation. We adopted a multi-stage strategy for improving the quality of registration. We also developed a visualisation tool to view registered pairs of WSIs at different magnifications. With the help of this tool, one can apply a transformation on the fly without the need to generate transformed source WSI in a pyramidal form. We compared the performance of data-driven features with that of hand-crafted features on the COMET dataset. Our approach can align the images with low registration errors. Generally, the success of non-rigid registration is dependent on the quality of rigid registration. To evaluate the efficacy of the DFBR method, the first two steps of the ANHIR winner's framework are replaced with our DFBR to register challenge provided image pairs. The modified framework produce comparable results to that of challenge winning team.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge