CliNER 2.0: Accessible and Accurate Clinical Concept Extraction
Paper and Code
Mar 06, 2018
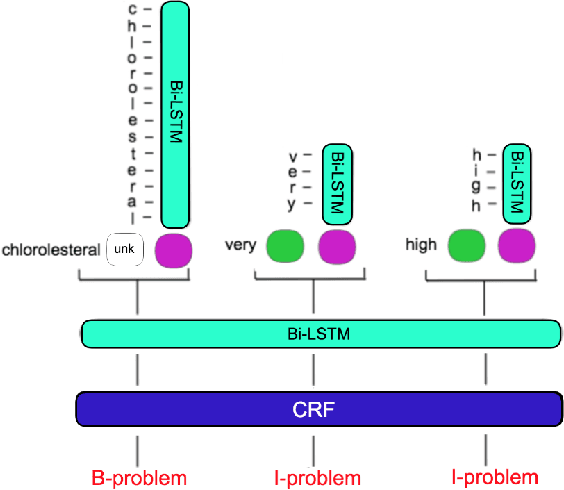

Clinical notes often describe important aspects of a patient's stay and are therefore critical to medical research. Clinical concept extraction (CCE) of named entities - such as problems, tests, and treatments - aids in forming an understanding of notes and provides a foundation for many downstream clinical decision-making tasks. Historically, this task has been posed as a standard named entity recognition (NER) sequence tagging problem, and solved with feature-based methods using handengineered domain knowledge. Recent advances, however, have demonstrated the efficacy of LSTM-based models for NER tasks, including CCE. This work presents CliNER 2.0, a simple-to-install, open-source tool for extracting concepts from clinical text. CliNER 2.0 uses a word- and character- level LSTM model, and achieves state-of-the-art performance. For ease of use, the tool also includes pre-trained models available for public use.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge