Chemical-induced Disease Relation Extraction with Dependency Information and Prior Knowledge
Paper and Code
Jan 02, 2020
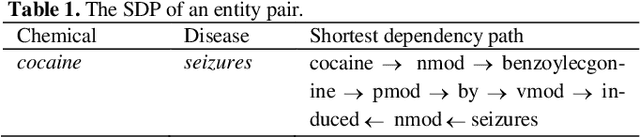
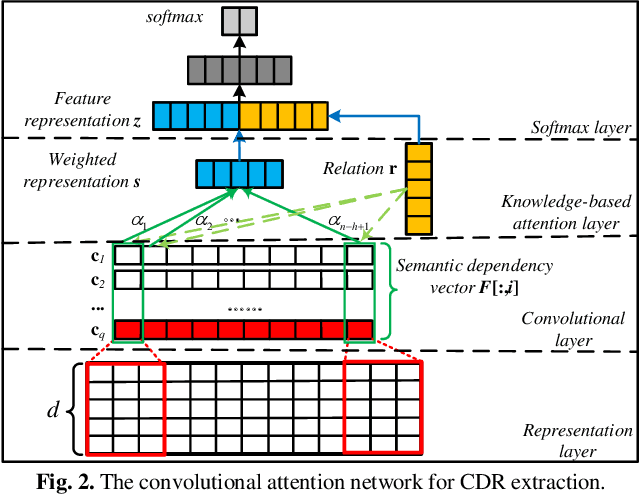
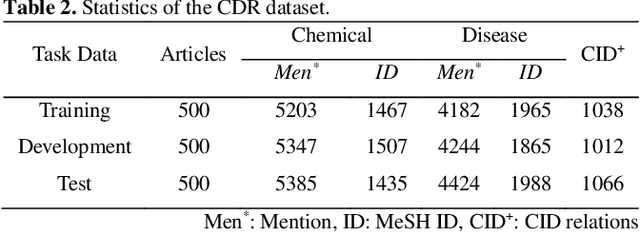
Chemical-disease relation (CDR) extraction is significantly important to various areas of biomedical research and health care. Nowadays, many large-scale biomedical knowledge bases (KBs) containing triples about entity pairs and their relations have been built. KBs are important resources for biomedical relation extraction. However, previous research pays little attention to prior knowledge. In addition, the dependency tree contains important syntactic and semantic information, which helps to improve relation extraction. So how to effectively use it is also worth studying. In this paper, we propose a novel convolutional attention network (CAN) for CDR extraction. Firstly, we extract the shortest dependency path (SDP) between chemical and disease pairs in a sentence, which includes a sequence of words, dependency directions, and dependency relation tags. Then the convolution operations are performed on the SDP to produce deep semantic dependency features. After that, an attention mechanism is employed to learn the importance/weight of each semantic dependency vector related to knowledge representations learned from KBs. Finally, in order to combine dependency information and prior knowledge, the concatenation of weighted semantic dependency representations and knowledge representations is fed to the softmax layer for classification. Experiments on the BioCreative V CDR dataset show that our method achieves comparable performance with the state-of-the-art systems, and both dependency information and prior knowledge play important roles in CDR extraction task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge