Brain dynamics via Cumulative Auto-Regressive Self-Attention
Paper and Code
Nov 01, 2021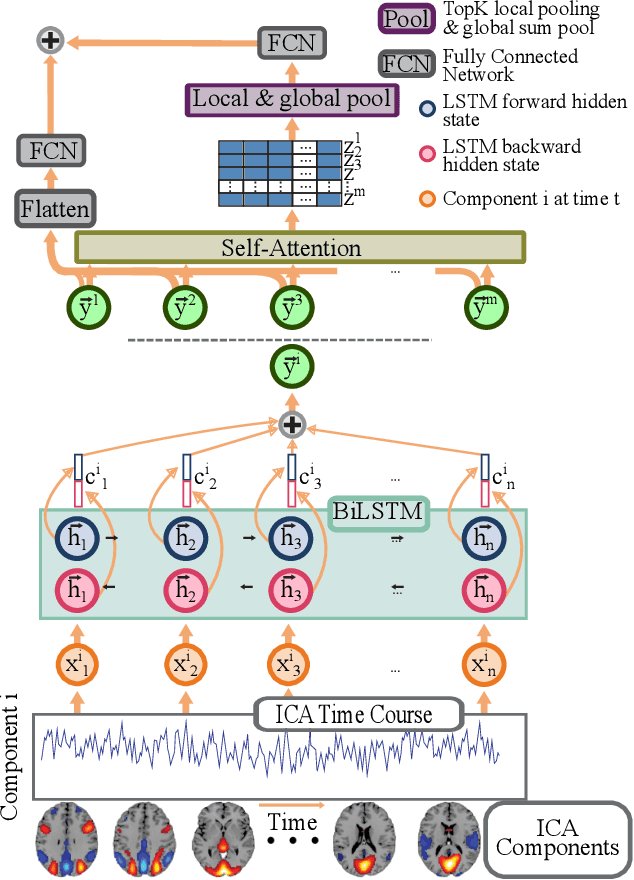

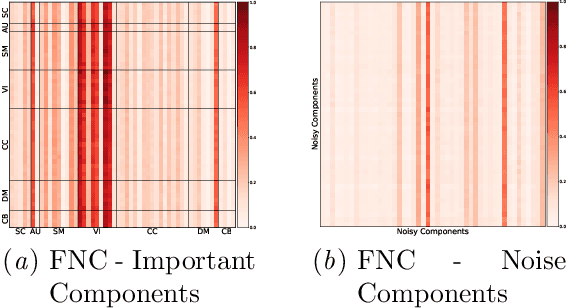
Multivariate dynamical processes can often be intuitively described by a weighted connectivity graph between components representing each individual time-series. Even a simple representation of this graph as a Pearson correlation matrix may be informative and predictive as demonstrated in the brain imaging literature. However, there is a consensus expectation that powerful graph neural networks (GNNs) should perform better in similar settings. In this work, we present a model that is considerably shallow than deep GNNs, yet outperforms them in predictive accuracy in a brain imaging application. Our model learns the autoregressive structure of individual time series and estimates directed connectivity graphs between the learned representations via a self-attention mechanism in an end-to-end fashion. The supervised training of the model as a classifier between patients and controls results in a model that generates directed connectivity graphs and highlights the components of the time-series that are predictive for each subject. We demonstrate our results on a functional neuroimaging dataset classifying schizophrenia patients and controls.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge