BMapOpt: Optimization of Brain Tissue Probability Maps using a Differentiable MRI Simulator
Paper and Code
Apr 23, 2024
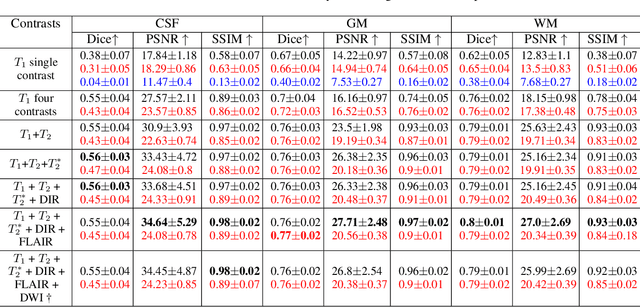
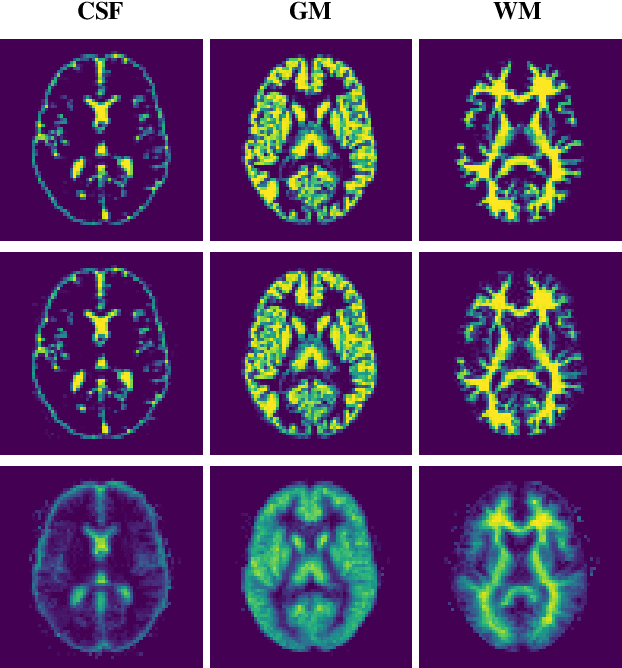
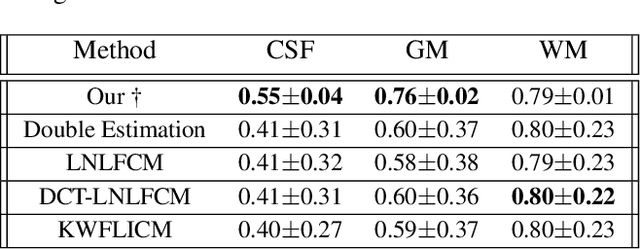
Reconstructing digital brain phantoms in the form of multi-channeled brain tissue probability maps for individual subjects is essential for capturing brain anatomical variability, understanding neurological diseases, as well as for testing image processing methods. We demonstrate the first framework that optimizes brain tissue probability maps (Gray Matter - GM, White Matter - WM, and Cerebrospinal fluid - CSF) with the help of a Physics-based differentiable MRI simulator that models the magnetization signal at each voxel in the image. Given an observed $T_1$/$T_2$-weighted MRI scan, the corresponding clinical MRI sequence, and the MRI differentiable simulator, we optimize the simulator's input probability maps by back-propagating the L2 loss between the simulator's output and the $T_1$/$T_2$-weighted scan. This approach has the significant advantage of not relying on any training data, and instead uses the strong inductive bias of the MRI simulator. We tested the model on 20 scans from the BrainWeb database and demonstrate a highly accurate reconstruction of GM, WM, and CSF.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge