Autoencoders for Unsupervised Anomaly Segmentation in Brain MR Images: A Comparative Study
Paper and Code
Apr 08, 2020
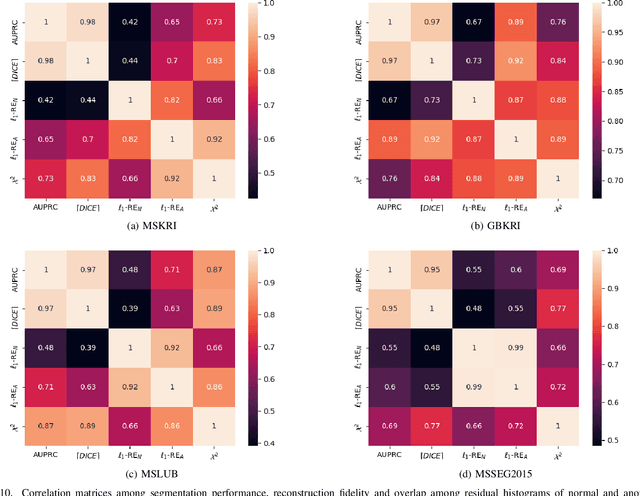
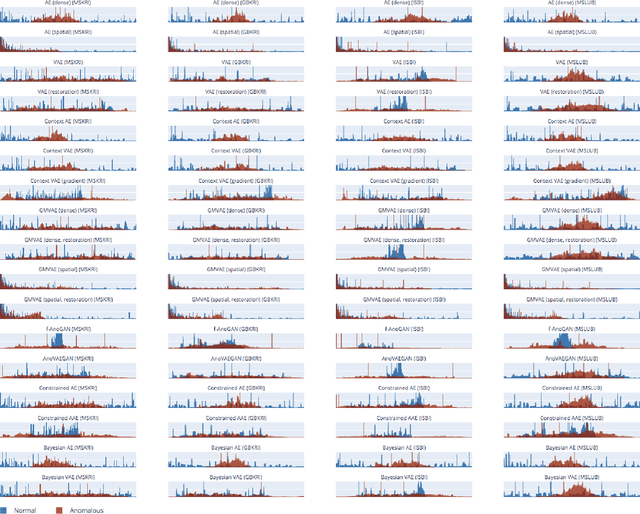
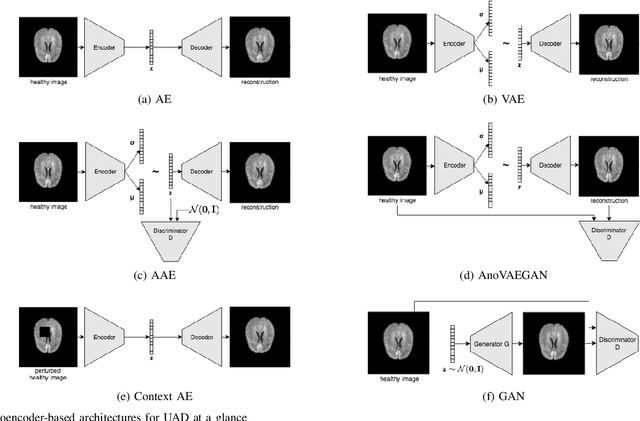
Deep unsupervised representation learning has recently led to new approaches in the field of Unsupervised Anomaly Detection (UAD) in brain MRI. The main principle behind these works is to learn a model of normal anatomy by learning to compress and recover healthy data. This allows to spot abnormal structures from erroneous recoveries of compressed, potentially anomalous samples. The concept is of great interest to the medical image analysis community as it i) relieves from the need of vast amounts of manually segmented training data---a necessity for and pitfall of current supervised Deep Learning---and ii) theoretically allows to detect arbitrary, even rare pathologies which supervised approaches might fail to find. To date, the experimental design of most works hinders a valid comparison, because i) they are evaluated against different datasets and different pathologies, ii) use different image resolutions and iii) different model architectures with varying complexity. The intent of this work is to establish comparability among recent methods by utilizing a single architecture, a single resolution and the same dataset(s). Besides providing a ranking of the methods, we also try to answer questions like i) how many healthy training subjects are needed to model normality and ii) if the reviewed approaches are also sensitive to domain shift. Further, we identify open challenges and provide suggestions for future community efforts and research directions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge