Zanting Ye
Unveiling and Bridging the Functional Perception Gap in MLLMs: Atomic Visual Alignment and Hierarchical Evaluation via PET-Bench
Jan 06, 2026Abstract:While Multimodal Large Language Models (MLLMs) have demonstrated remarkable proficiency in tasks such as abnormality detection and report generation for anatomical modalities, their capability in functional imaging remains largely unexplored. In this work, we identify and quantify a fundamental functional perception gap: the inability of current vision encoders to decode functional tracer biodistribution independent of morphological priors. Identifying Positron Emission Tomography (PET) as the quintessential modality to investigate this disconnect, we introduce PET-Bench, the first large-scale functional imaging benchmark comprising 52,308 hierarchical QA pairs from 9,732 multi-site, multi-tracer PET studies. Extensive evaluation of 19 state-of-the-art MLLMs reveals a critical safety hazard termed the Chain-of-Thought (CoT) hallucination trap. We observe that standard CoT prompting, widely considered to enhance reasoning, paradoxically decouples linguistic generation from visual evidence in PET, producing clinically fluent but factually ungrounded diagnoses. To resolve this, we propose Atomic Visual Alignment (AVA), a simple fine-tuning strategy that enforces the mastery of low-level functional perception prior to high-level diagnostic reasoning. Our results demonstrate that AVA effectively bridges the perception gap, transforming CoT from a source of hallucination into a robust inference tool and improving diagnostic accuracy by up to 14.83%. Code and data are available at https://github.com/yezanting/PET-Bench.
MDAA-Diff: CT-Guided Multi-Dose Adaptive Attention Diffusion Model for PET Denoising
May 08, 2025Abstract:Acquiring high-quality Positron Emission Tomography (PET) images requires administering high-dose radiotracers, which increases radiation exposure risks. Generating standard-dose PET (SPET) from low-dose PET (LPET) has become a potential solution. However, previous studies have primarily focused on single low-dose PET denoising, neglecting two critical factors: discrepancies in dose response caused by inter-patient variability, and complementary anatomical constraints derived from CT images. In this work, we propose a novel CT-Guided Multi-dose Adaptive Attention Denoising Diffusion Model (MDAA-Diff) for multi-dose PET denoising. Our approach integrates anatomical guidance and dose-level adaptation to achieve superior denoising performance under low-dose conditions. Specifically, this approach incorporates a CT-Guided High-frequency Wavelet Attention (HWA) module, which uses wavelet transforms to separate high-frequency anatomical boundary features from CT images. These extracted features are then incorporated into PET imaging through an adaptive weighted fusion mechanism to enhance edge details. Additionally, we propose the Dose-Adaptive Attention (DAA) module, a dose-conditioned enhancement mechanism that dynamically integrates dose levels into channel-spatial attention weight calculation. Extensive experiments on 18F-FDG and 68Ga-FAPI datasets demonstrate that MDAA-Diff outperforms state-of-the-art approaches in preserving diagnostic quality under reduced-dose conditions. Our code is publicly available.
Semi-KAN: KAN Provides an Effective Representation for Semi-Supervised Learning in Medical Image Segmentation
Mar 19, 2025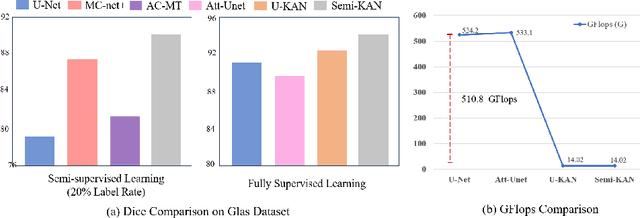
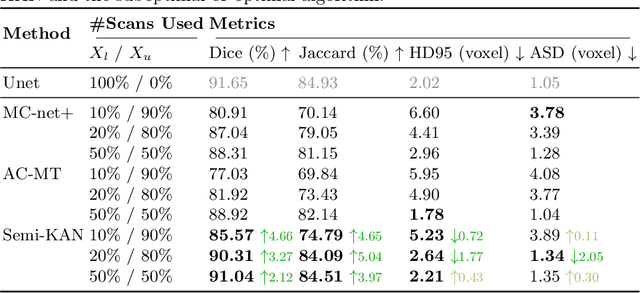
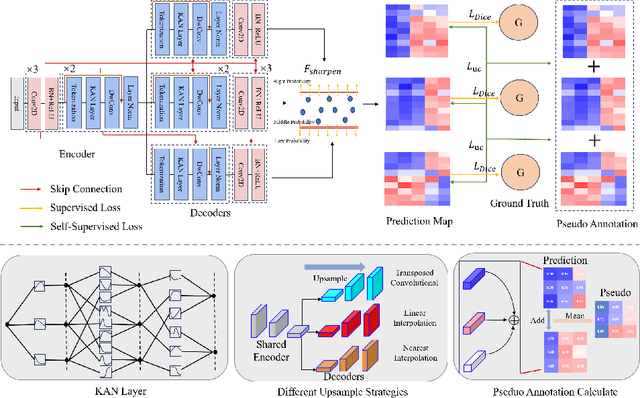
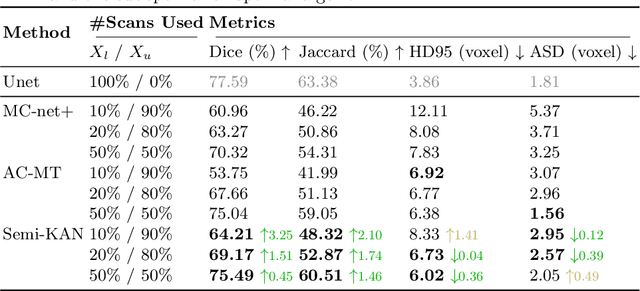
Abstract:Deep learning-based medical image segmentation has shown remarkable success; however, it typically requires extensive pixel-level annotations, which are both expensive and time-intensive. Semi-supervised medical image segmentation (SSMIS) offers a viable alternative, driven by advancements in CNNs and ViTs. However, these networks often rely on single fixed activation functions and linear modeling patterns, limiting their ability to effectively learn robust representations. Given the limited availability of labeled date, achieving robust representation learning becomes crucial. Inspired by Kolmogorov-Arnold Networks (KANs), we propose Semi-KAN, which leverages the untapped potential of KANs to enhance backbone architectures for representation learning in SSMIS. Our findings indicate that: (1) compared to networks with fixed activation functions, KANs exhibit superior representation learning capabilities with fewer parameters, and (2) KANs excel in high-semantic feature spaces. Building on these insights, we integrate KANs into tokenized intermediate representations, applying them selectively at the encoder's bottleneck and the decoder's top layers within a U-Net pipeline to extract high-level semantic features. Although learnable activation functions improve feature expansion, they introduce significant computational overhead with only marginal performance gains. To mitigate this, we reduce the feature dimensions and employ horizontal scaling to capture multiple pattern representations. Furthermore, we design a multi-branch U-Net architecture with uncertainty estimation to effectively learn diverse pattern representations. Extensive experiments on four public datasets demonstrate that Semi-KAN surpasses baseline networks, utilizing fewer KAN layers and lower computational cost, thereby underscoring the potential of KANs as a promising approach for SSMIS.
Self is the Best Learner: CT-free Ultra-Low-Dose PET Organ Segmentation via Collaborating Denoising and Segmentation Learning
Mar 05, 2025Abstract:Organ segmentation in Positron Emission Tomography (PET) plays a vital role in cancer quantification. Low-dose PET (LDPET) provides a safer alternative by reducing radiation exposure. However, the inherent noise and blurred boundaries make organ segmentation more challenging. Additionally, existing PET organ segmentation methods rely on co-registered Computed Tomography (CT) annotations, overlooking the problem of modality mismatch. In this study, we propose LDOS, a novel CT-free ultra-LDPET organ segmentation pipeline. Inspired by Masked Autoencoders (MAE), we reinterpret LDPET as a naturally masked version of Full-Dose PET (FDPET). LDOS adopts a simple yet effective architecture: a shared encoder extracts generalized features, while task-specific decoders independently refine outputs for denoising and segmentation. By integrating CT-derived organ annotations into the denoising process, LDOS improves anatomical boundary recognition and alleviates the PET/CT misalignments. Experiments demonstrate that LDOS achieves state-of-the-art performance with mean Dice scores of 73.11% (18F-FDG) and 73.97% (68Ga-FAPI) across 18 organs in 5% dose PET. Our code is publicly available.
SSL-DG: Rethinking and Fusing Semi-supervised Learning and Domain Generalization in Medical Image Segmentation
Nov 05, 2023Abstract:Deep learning-based medical image segmentation is an essential yet challenging task in clinical practice, which arises from restricted access to annotated data coupled with the occurrence of domain shifts. Previous attempts have focused on isolated solutions, while disregarding their inter-connectedness. In this paper, we rethink the relationship between semi-supervised learning (SSL) and domain generalization (DG), which are the cutting-edge approaches to address the annotated data-driven constraints and the domain shift issues. Inspired by class-level representation, we show that unseen target data can be represented by a linear combination of source data, which can be achieved by simple data augmentation. The augmented data enrich domain distributions while having semantic consistency, aligning with the principles of consistency-based SSL. Accordingly, we propose SSL-DG, fusing DG and SSL, to achieve cross-domain generalization with limited annotations. Specifically, the global and focal region augmentation, together with an augmentation scale-balancing mechanism, are used to construct a mask-based domain diffusion augmentation module to significantly enrich domain diversity. In order to obtain consistent predictions for the same source data in different networks, we use uncertainty estimation and a deep mutual learning strategy to enforce the consistent constraint. Extensive experiments including ablation studies are designed to validate the proposed SSL-DG. The results demonstrate that our SSL-DG significantly outperforms state-of-the-art solutions in two challenging DG tasks with limited annotations. Code is available at https://github.com/yezanting/SSL-DG.
MLN-net: A multi-source medical image segmentation method for clustered microcalcifications using multiple layer normalization
Sep 06, 2023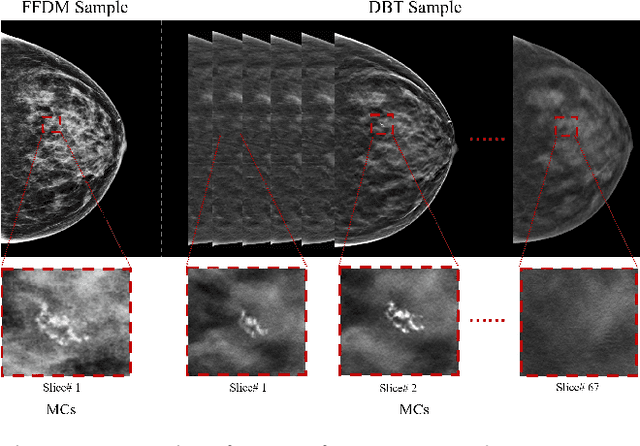


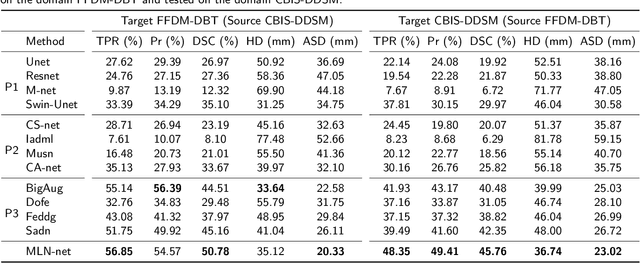
Abstract:Accurate segmentation of clustered microcalcifications in mammography is crucial for the diagnosis and treatment of breast cancer. Despite exhibiting expert-level accuracy, recent deep learning advancements in medical image segmentation provide insufficient contribution to practical applications, due to the domain shift resulting from differences in patient postures, individual gland density, and imaging modalities of mammography etc. In this paper, a novel framework named MLN-net, which can accurately segment multi-source images using only single source images, is proposed for clustered microcalcification segmentation. We first propose a source domain image augmentation method to generate multi-source images, leading to improved generalization. And a structure of multiple layer normalization (LN) layers is used to construct the segmentation network, which can be found efficient for clustered microcalcification segmentation in different domains. Additionally, a branch selection strategy is designed for measuring the similarity of the source domain data and the target domain data. To validate the proposed MLN-net, extensive analyses including ablation experiments are performed, comparison of 12 baseline methods. Extensive experiments validate the effectiveness of MLN-net in segmenting clustered microcalcifications from different domains and the its segmentation accuracy surpasses state-of-the-art methods. Code will be available at https://github.com/yezanting/MLN-NET-VERSON1.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge