Won-Ki Jeong
Zero-shot Volumetric CT Super-Resolution using 3D Gaussian Splatting with Upsampled 2D X-ray Projection Priors
Aug 21, 2025Abstract:Computed tomography (CT) is widely used in clinical diagnosis, but acquiring high-resolution (HR) CT is limited by radiation exposure risks. Deep learning-based super-resolution (SR) methods have been studied to reconstruct HR from low-resolution (LR) inputs. While supervised SR approaches have shown promising results, they require large-scale paired LR-HR volume datasets that are often unavailable. In contrast, zero-shot methods alleviate the need for paired data by using only a single LR input, but typically struggle to recover fine anatomical details due to limited internal information. To overcome these, we propose a novel zero-shot 3D CT SR framework that leverages upsampled 2D X-ray projection priors generated by a diffusion model. Exploiting the abundance of HR 2D X-ray data, we train a diffusion model on large-scale 2D X-ray projection and introduce a per-projection adaptive sampling strategy. It selects the generative process for each projection, thus providing HR projections as strong external priors for 3D CT reconstruction. These projections serve as inputs to 3D Gaussian splatting for reconstructing a 3D CT volume. Furthermore, we propose negative alpha blending (NAB-GS) that allows negative values in Gaussian density representation. NAB-GS enables residual learning between LR and diffusion-based projections, thereby enhancing high-frequency structure reconstruction. Experiments on two datasets show that our method achieves superior quantitative and qualitative results for 3D CT SR.
WISE-FUSE: Efficient Whole Slide Image Encoding via Coarse-to-Fine Patch Selection with VLM and LLM Knowledge Fusion
Aug 20, 2025



Abstract:Whole slide images (WSIs) in computational pathology (CPath) pose a major computational challenge due to their gigapixel scale, often requiring the processing of tens to hundreds of thousands of high-resolution patches per slide. This results in prohibitive encoding costs, with preprocessing and training times extending to days or even weeks-making WSI encoding the most significant bottleneck in real-world deployment. In this work, we propose WISE-FUSE, an adaptive WSI encoding framework that leverages pathology-domain vision-language models and large language models to address this challenge by selectively processing diagnostically relevant regions. WISE-FUSE first computes similarity scores between low-resolution patches and class-specific textual descriptions using a knowledge distillation mechanism that preserves fine-grained diagnostic features. Based on these similarity scores, we select a small subset of informative regions for the target task, which quickly eliminates irrelevant patches at the coarse level. The corresponding high-resolution patches are then selectively encoded and fused with textual embeddings to reinforce diagnostic context. Extensive experiments demonstrate that WISE-FUSE reduces WSI encoding time by over threefold while achieving diagnostic performance comparable to or surpassing that of exhaustive patch processing, offering a scalable and practical solution for CPath.
Virtual Multiplex Staining for Histological Images using a Marker-wise Conditioned Diffusion Model
Aug 20, 2025Abstract:Multiplex imaging is revolutionizing pathology by enabling the simultaneous visualization of multiple biomarkers within tissue samples, providing molecular-level insights that traditional hematoxylin and eosin (H&E) staining cannot provide. However, the complexity and cost of multiplex data acquisition have hindered its widespread adoption. Additionally, most existing large repositories of H&E images lack corresponding multiplex images, limiting opportunities for multimodal analysis. To address these challenges, we leverage recent advances in latent diffusion models (LDMs), which excel at modeling complex data distributions utilizing their powerful priors for fine-tuning to a target domain. In this paper, we introduce a novel framework for virtual multiplex staining that utilizes pretrained LDM parameters to generate multiplex images from H&E images using a conditional diffusion model. Our approach enables marker-by-marker generation by conditioning the diffusion model on each marker, while sharing the same architecture across all markers. To tackle the challenge of varying pixel value distributions across different marker stains and to improve inference speed, we fine-tune the model for single-step sampling, enhancing both color contrast fidelity and inference efficiency through pixel-level loss functions. We validate our framework on two publicly available datasets, notably demonstrating its effectiveness in generating up to 18 different marker types with improved accuracy, a substantial increase over the 2-3 marker types achieved in previous approaches. This validation highlights the potential of our framework, pioneering virtual multiplex staining. Finally, this paper bridges the gap between H&E and multiplex imaging, potentially enabling retrospective studies and large-scale analyses of existing H&E image repositories.
MC-INR: Efficient Encoding of Multivariate Scientific Simulation Data using Meta-Learning and Clustered Implicit Neural Representations
Jul 03, 2025Abstract:Implicit Neural Representations (INRs) are widely used to encode data as continuous functions, enabling the visualization of large-scale multivariate scientific simulation data with reduced memory usage. However, existing INR-based methods face three main limitations: (1) inflexible representation of complex structures, (2) primarily focusing on single-variable data, and (3) dependence on structured grids. Thus, their performance degrades when applied to complex real-world datasets. To address these limitations, we propose a novel neural network-based framework, MC-INR, which handles multivariate data on unstructured grids. It combines meta-learning and clustering to enable flexible encoding of complex structures. To further improve performance, we introduce a residual-based dynamic re-clustering mechanism that adaptively partitions clusters based on local error. We also propose a branched layer to leverage multivariate data through independent branches simultaneously. Experimental results demonstrate that MC-INR outperforms existing methods on scientific data encoding tasks.
Reference-free Axial Super-resolution of 3D Microscopy Images using Implicit Neural Representation with a 2D Diffusion Prior
Aug 16, 2024Abstract:Analysis and visualization of 3D microscopy images pose challenges due to anisotropic axial resolution, demanding volumetric super-resolution along the axial direction. While training a learning-based 3D super-resolution model seems to be a straightforward solution, it requires ground truth isotropic volumes and suffers from the curse of dimensionality. Therefore, existing methods utilize 2D neural networks to reconstruct each axial slice, eventually piecing together the entire volume. However, reconstructing each slice in the pixel domain fails to give consistent reconstruction in all directions leading to misalignment artifacts. In this work, we present a reconstruction framework based on implicit neural representation (INR), which allows 3D coherency even when optimized by independent axial slices in a batch-wise manner. Our method optimizes a continuous volumetric representation from low-resolution axial slices, using a 2D diffusion prior trained on high-resolution lateral slices without requiring isotropic volumes. Through experiments on real and synthetic anisotropic microscopy images, we demonstrate that our method surpasses other state-of-the-art reconstruction methods. The source code is available on GitHub: https://github.com/hvcl/INR-diffusion.
Controllable and Efficient Multi-Class Pathology Nuclei Data Augmentation using Text-Conditioned Diffusion Models
Jul 19, 2024



Abstract:In the field of computational pathology, deep learning algorithms have made significant progress in tasks such as nuclei segmentation and classification. However, the potential of these advanced methods is limited by the lack of available labeled data. Although image synthesis via recent generative models has been actively explored to address this challenge, existing works have barely addressed label augmentation and are mostly limited to single-class and unconditional label generation. In this paper, we introduce a novel two-stage framework for multi-class nuclei data augmentation using text-conditional diffusion models. In the first stage, we innovate nuclei label synthesis by generating multi-class semantic labels and corresponding instance maps through a joint diffusion model conditioned by text prompts that specify the label structure information. In the second stage, we utilize a semantic and text-conditional latent diffusion model to efficiently generate high-quality pathology images that align with the generated nuclei label images. We demonstrate the effectiveness of our method on large and diverse pathology nuclei datasets, with evaluations including qualitative and quantitative analyses, as well as assessments of downstream tasks.
Co-synthesis of Histopathology Nuclei Image-Label Pairs using a Context-Conditioned Joint Diffusion Model
Jul 19, 2024Abstract:In multi-class histopathology nuclei analysis tasks, the lack of training data becomes a main bottleneck for the performance of learning-based methods. To tackle this challenge, previous methods have utilized generative models to increase data by generating synthetic samples. However, existing methods often overlook the importance of considering the context of biological tissues (e.g., shape, spatial layout, and tissue type) in the synthetic data. Moreover, while generative models have shown superior performance in synthesizing realistic histopathology images, none of the existing methods are capable of producing image-label pairs at the same time. In this paper, we introduce a novel framework for co-synthesizing histopathology nuclei images and paired semantic labels using a context-conditioned joint diffusion model. We propose conditioning of a diffusion model using nucleus centroid layouts with structure-related text prompts to incorporate spatial and structural context information into the generation targets. Moreover, we enhance the granularity of our synthesized semantic labels by generating instance-wise nuclei labels using distance maps synthesized concurrently in conjunction with the images and semantic labels. We demonstrate the effectiveness of our framework in generating high-quality samples on multi-institutional, multi-organ, and multi-modality datasets. Our synthetic data consistently outperforms existing augmentation methods in the downstream tasks of nuclei segmentation and classification.
Evaluation and improvement of Segment Anything Model for interactive histopathology image segmentation
Oct 16, 2023



Abstract:With the emergence of the Segment Anything Model (SAM) as a foundational model for image segmentation, its application has been extensively studied across various domains, including the medical field. However, its potential in the context of histopathology data, specifically in region segmentation, has received relatively limited attention. In this paper, we evaluate SAM's performance in zero-shot and fine-tuned scenarios on histopathology data, with a focus on interactive segmentation. Additionally, we compare SAM with other state-of-the-art interactive models to assess its practical potential and evaluate its generalization capability with domain adaptability. In the experimental results, SAM exhibits a weakness in segmentation performance compared to other models while demonstrating relative strengths in terms of inference time and generalization capability. To improve SAM's limited local refinement ability and to enhance prompt stability while preserving its core strengths, we propose a modification of SAM's decoder. The experimental results suggest that the proposed modification is effective to make SAM useful for interactive histology image segmentation. The code is available at \url{https://github.com/hvcl/SAM_Interactive_Histopathology}
Reference-Free Isotropic 3D EM Reconstruction using Diffusion Models
Aug 03, 2023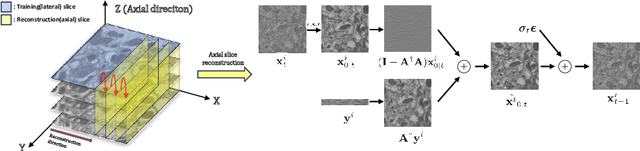
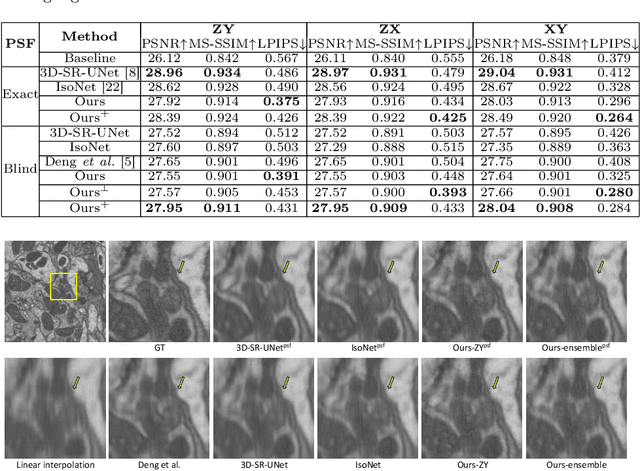
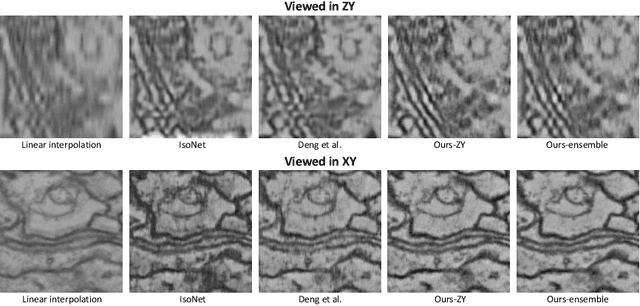
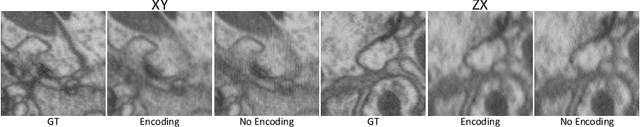
Abstract:Electron microscopy (EM) images exhibit anisotropic axial resolution due to the characteristics inherent to the imaging modality, presenting challenges in analysis and downstream tasks.In this paper, we propose a diffusion-model-based framework that overcomes the limitations of requiring reference data or prior knowledge about the degradation process. Our approach utilizes 2D diffusion models to consistently reconstruct 3D volumes and is well-suited for highly downsampled data. Extensive experiments conducted on two public datasets demonstrate the robustness and superiority of leveraging the generative prior compared to supervised learning methods. Additionally, we demonstrate our method's feasibility for self-supervised reconstruction, which can restore a single anisotropic volume without any training data.
CGAM: Click-Guided Attention Module for Interactive Pathology Image Segmentation via Backpropagating Refinement
Jul 03, 2023



Abstract:Tumor region segmentation is an essential task for the quantitative analysis of digital pathology. Recently presented deep neural networks have shown state-of-the-art performance in various image-segmentation tasks. However, because of the unclear boundary between the cancerous and normal regions in pathology images, despite using modern methods, it is difficult to produce satisfactory segmentation results in terms of the reliability and accuracy required for medical data. In this study, we propose an interactive segmentation method that allows users to refine the output of deep neural networks through click-type user interactions. The primary method is to formulate interactive segmentation as an optimization problem that leverages both user-provided click constraints and semantic information in a feature map using a click-guided attention module (CGAM). Unlike other existing methods, CGAM avoids excessive changes in segmentation results, which can lead to the overfitting of user clicks. Another advantage of CGAM is that the model size is independent of input image size. Experimental results on pathology image datasets indicated that our method performs better than existing state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge