Till Huelnhagen
Fast refacing of MR images with a generative neural network lowers re-identification risk and preserves volumetric consistency
May 26, 2023Abstract:With the rise of open data, identifiability of individuals based on 3D renderings obtained from routine structural magnetic resonance imaging (MRI) scans of the head has become a growing privacy concern. To protect subject privacy, several algorithms have been developed to de-identify imaging data using blurring, defacing or refacing. Completely removing facial structures provides the best re-identification protection but can significantly impact post-processing steps, like brain morphometry. As an alternative, refacing methods that replace individual facial structures with generic templates have a lower effect on the geometry and intensity distribution of original scans, and are able to provide more consistent post-processing results by the price of higher re-identification risk and computational complexity. In the current study, we propose a novel method for anonymised face generation for defaced 3D T1-weighted scans based on a 3D conditional generative adversarial network. To evaluate the performance of the proposed de-identification tool, a comparative study was conducted between several existing defacing and refacing tools, with two different segmentation algorithms (FAST and Morphobox). The aim was to evaluate (i) impact on brain morphometry reproducibility, (ii) re-identification risk, (iii) balance between (i) and (ii), and (iv) the processing time. The proposed method takes 9 seconds for face generation and is suitable for recovering consistent post-processing results after defacing.
Cortical lesions, central vein sign, and paramagnetic rim lesions in multiple sclerosis: emerging machine learning techniques and future avenues
Jan 19, 2022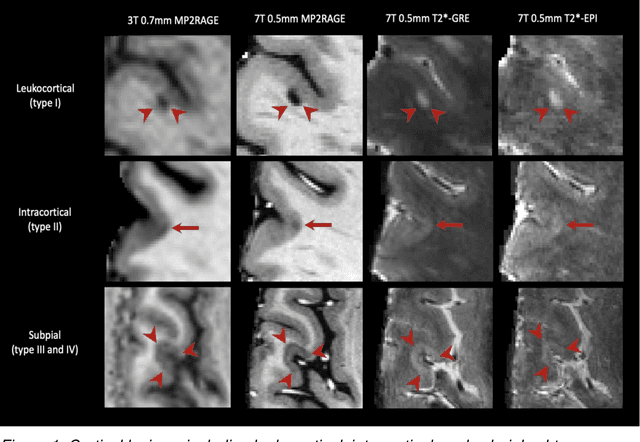
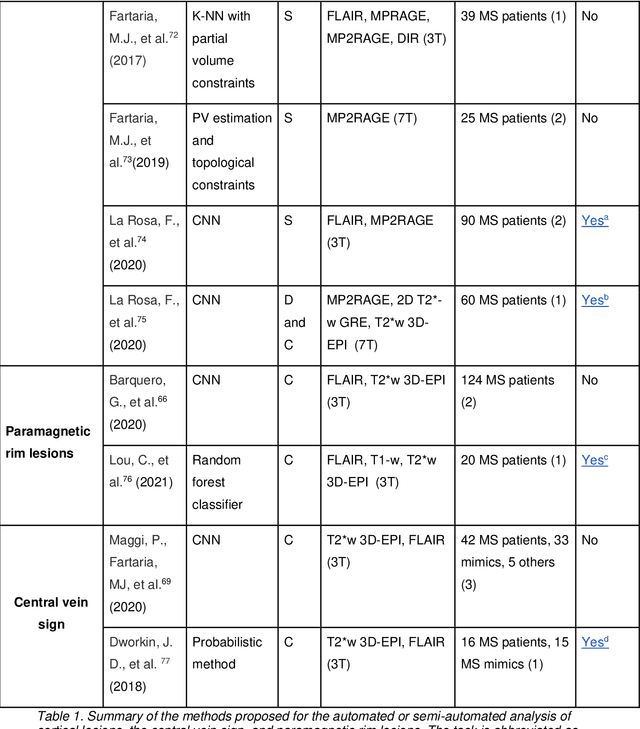
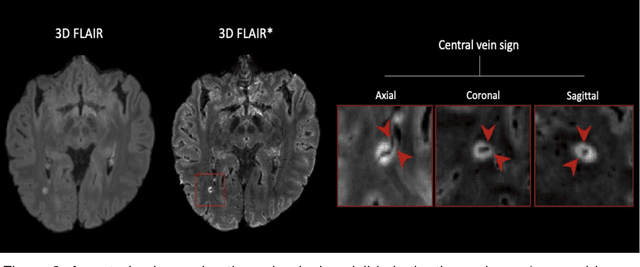
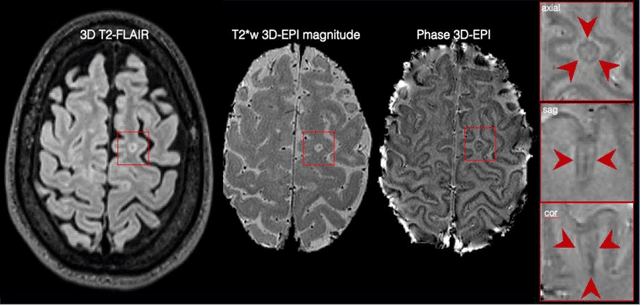
Abstract:The current multiple sclerosis (MS) diagnostic criteria lack specificity, and this may lead to misdiagnosis, which remains an issue in present-day clinical practice. In addition, conventional biomarkers only moderately correlate with MS disease progression. Recently, advanced MS lesional imaging biomarkers such as cortical lesions (CL), the central vein sign (CVS), and paramagnetic rim lesions (PRL), visible in specialized magnetic resonance imaging (MRI) sequences, have shown higher specificity in differential diagnosis. Moreover, studies have shown that CL and PRL are potential prognostic biomarkers, the former correlating with cognitive impairments and the latter with early disability progression. As machine learning-based methods have achieved extraordinary performance in the assessment of conventional imaging biomarkers, such as white matter lesion segmentation, several automated or semi-automated methods have been proposed for CL, CVS, and PRL as well. In the present review, we first introduce these advanced MS imaging biomarkers and their imaging methods. Subsequently, we describe the corresponding machine learning-based methods that were used to tackle these clinical questions, putting them into context with respect to the challenges they are still facing, including non-standardized MRI protocols, limited datasets, and moderate inter-rater variability. We conclude by presenting the current limitations that prevent their broader deployment and suggesting future research directions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge