Shreyas Cholia
Disentangling multiple scattering with deep learning: application to strain mapping from electron diffraction patterns
Feb 01, 2022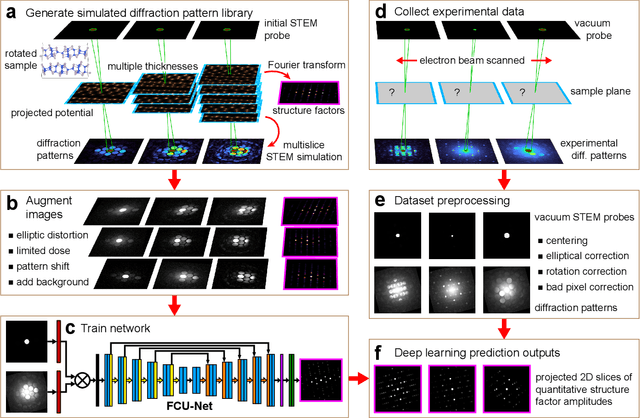
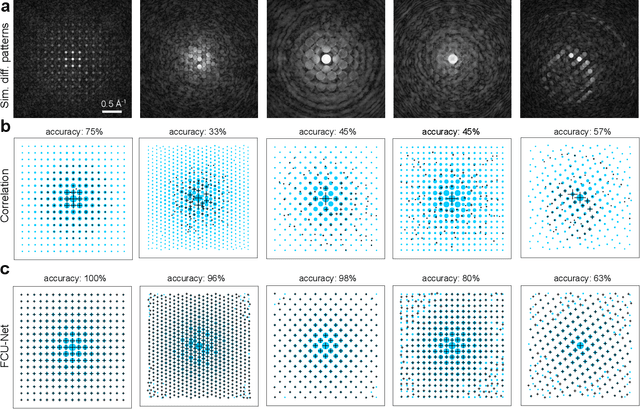
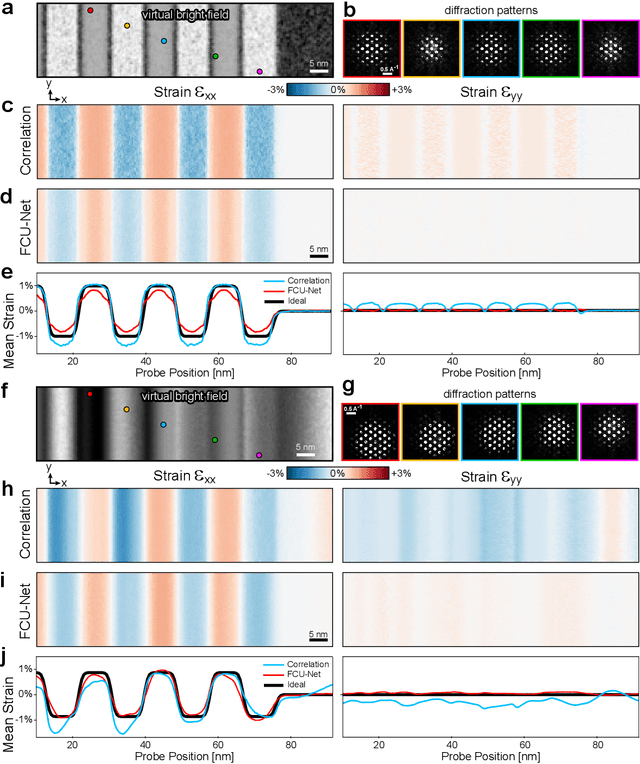

Abstract:Implementation of a fast, robust, and fully-automated pipeline for crystal structure determination and underlying strain mapping for crystalline materials is important for many technological applications. Scanning electron nanodiffraction offers a procedure for identifying and collecting strain maps with good accuracy and high spatial resolutions. However, the application of this technique is limited, particularly in thick samples where the electron beam can undergo multiple scattering, which introduces signal nonlinearities. Deep learning methods have the potential to invert these complex signals, but previous implementations are often trained only on specific crystal systems or a small subset of the crystal structure and microscope parameter phase space. In this study, we implement a Fourier space, complex-valued deep neural network called FCU-Net, to invert highly nonlinear electron diffraction patterns into the corresponding quantitative structure factor images. We trained the FCU-Net using over 200,000 unique simulated dynamical diffraction patterns which include many different combinations of crystal structures, orientations, thicknesses, microscope parameters, and common experimental artifacts. We evaluated the trained FCU-Net model against simulated and experimental 4D-STEM diffraction datasets, where it substantially out-performs conventional analysis methods. Our simulated diffraction pattern library, implementation of FCU-Net, and trained model weights are freely available in open source repositories, and can be adapted to many different diffraction measurement problems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge