Shalin B. Mehta
waveOrder: generalist framework for label-agnostic computational microscopy
Dec 13, 2024



Abstract:Correlative computational microscopy is accelerating the mapping of dynamic biological systems by integrating morphological and molecular measurements across spatial scales, from organelles to entire organisms. Visualization, measurement, and prediction of interactions among the components of biological systems can be accelerated by generalist computational imaging frameworks that relax the trade-offs imposed by multiplex dynamic imaging. This work reports a generalist framework for wave optical imaging of the architectural order (waveOrder) among biomolecules for encoding and decoding multiple specimen properties from a minimal set of acquired channels, with or without fluorescent labels. waveOrder expresses material properties in terms of elegant physically motivated basis vectors directly interpretable as phase, absorption, birefringence, diattenuation, and fluorophore density; and it expresses image data in terms of directly measurable Stokes parameters. We report a corresponding multi-channel reconstruction algorithm to recover specimen properties in multiple contrast modes. With this framework, we implement multiple 3D computational microscopy methods, including quantitative phase imaging, quantitative label-free imaging with phase and polarization, and fluorescence deconvolution imaging, across scales ranging from organelles to whole zebrafish. These advances are available via an extensible open-source computational imaging library, waveOrder, and a napari plugin, recOrder.
Contrastive learning of cell state dynamics in response to perturbations
Oct 15, 2024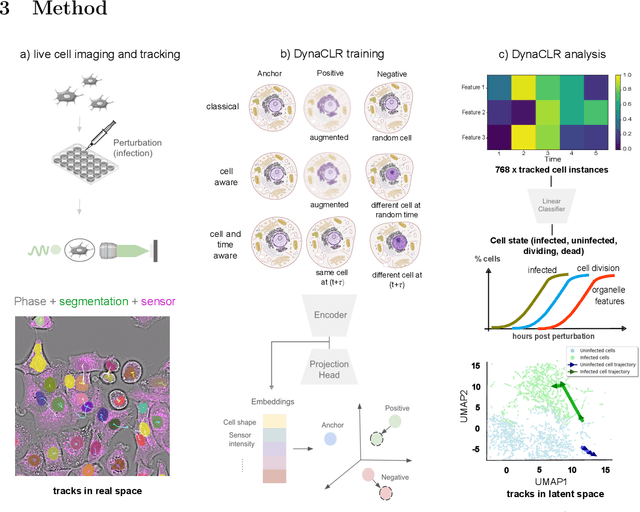
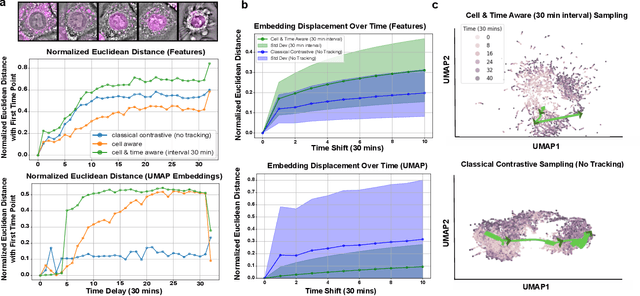
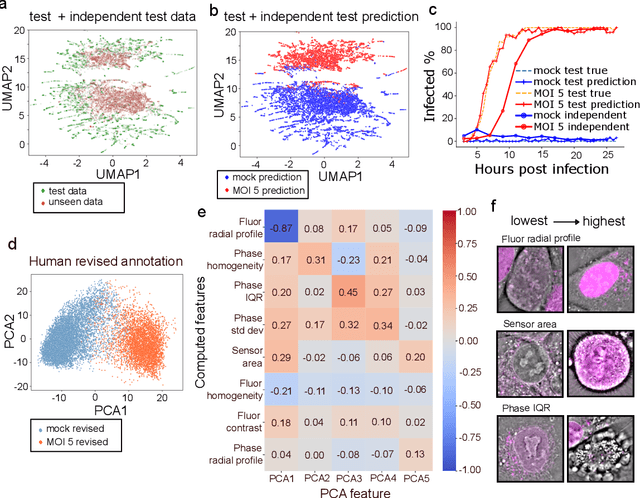
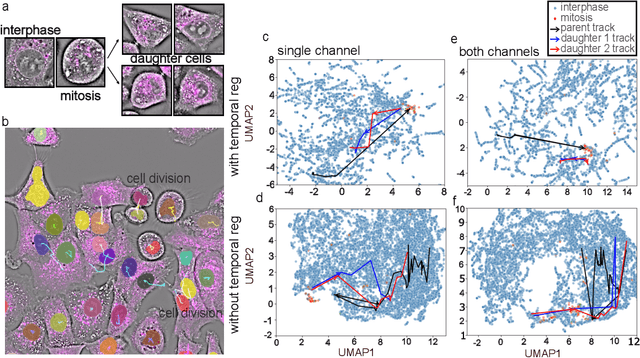
Abstract:We introduce DynaCLR, a self-supervised framework for modeling cell dynamics via contrastive learning of representations of time-lapse datasets. Live cell imaging of cells and organelles is widely used to analyze cellular responses to perturbations. Human annotation of dynamic cell states captured by time-lapse perturbation datasets is laborious and prone to bias. DynaCLR integrates single-cell tracking with time-aware contrastive learning to map images of cells at neighboring time points to neighboring embeddings. Mapping the morphological dynamics of cells to a temporally regularized embedding space makes the annotation, classification, clustering, or interpretation of the cell states more quantitative and efficient. We illustrate the features and applications of DynaCLR with the following experiments: analyzing the kinetics of viral infection in human cells, detecting transient changes in cell morphology due to cell division, and mapping the dynamics of organelles due to viral infection. Models trained with DynaCLR consistently achieve $>95\%$ accuracy for infection state classification, enable the detection of transient cell states and reliably embed unseen experiments. DynaCLR provides a flexible framework for comparative analysis of cell state dynamics due to perturbations, such as infection, gene knockouts, and drugs. We provide PyTorch-based implementations of the model training and inference pipeline (https://github.com/mehta-lab/viscy) and a user interface (https://github.com/czbiohub-sf/napari-iohub) for the visualization and annotation of trajectories of cells in the real space and the embedding space.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge