Poul Jørgen Jennum
Fully-automated sleep staging: multicenter validation of a generalizable deep neural network for Parkinson's disease and isolated REM sleep behavior disorder
Feb 11, 2026Abstract:Isolated REM sleep behavior disorder (iRBD) is a key prodromal marker of Parkinson's disease (PD), and video-polysomnography (vPSG) remains the diagnostic gold standard. However, manual sleep staging is particularly challenging in neurodegenerative diseases due to EEG abnormalities and fragmented sleep, making PSG assessments a bottleneck for deploying new RBD screening technologies at scale. We adapted U-Sleep, a deep neural network, for generalizable sleep staging in PD and iRBD. A pretrained U-Sleep model, based on a large, multisite non-neurodegenerative dataset (PUB; 19,236 PSGs across 12 sites), was fine-tuned on research datasets from two centers (Lundbeck Foundation Parkinson's Disease Research Center (PACE) and the Cologne-Bonn Cohort (CBC); 112 PD, 138 iRBD, 89 age-matched controls. The resulting model was evaluated on an independent dataset from the Danish Center for Sleep Medicine (DCSM; 81 PD, 36 iRBD, 87 sleep-clinic controls). A subset of PSGs with low agreement between the human rater and the model (Cohen's $κ$ < 0.6) was re-scored by a second blinded human rater to identify sources of disagreement. Finally, we applied confidence-based thresholds to optimize REM sleep staging. The pretrained model achieved mean $κ$ = 0.81 in PUB, but $κ$ = 0.66 when applied directly to PACE/CBC. By fine-tuning the model, we developed a generalized model with $κ$ = 0.74 on PACE/CBC (p < 0.001 vs. the pretrained model). In DCSM, mean and median $κ$ increased from 0.60 to 0.64 (p < 0.001) and 0.64 to 0.69 (p < 0.001), respectively. In the interrater study, PSGs with low agreement between the model and the initial scorer showed similarly low agreement between human scorers. Applying a confidence threshold increased the proportion of correctly identified REM sleep epochs from 85% to 95.5%, while preserving sufficient (> 5 min) REM sleep for 95% of subjects.
U-Time: A Fully Convolutional Network for Time Series Segmentation Applied to Sleep Staging
Oct 24, 2019
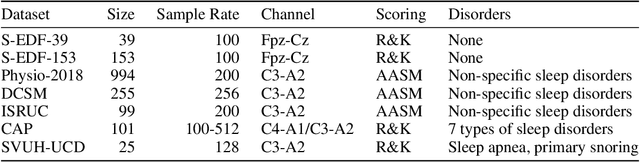
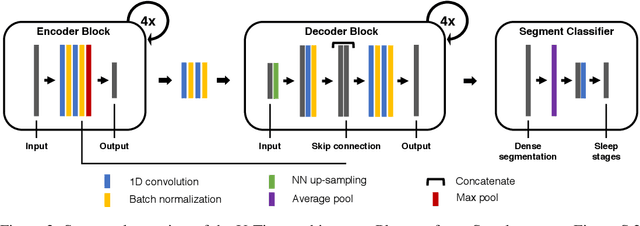
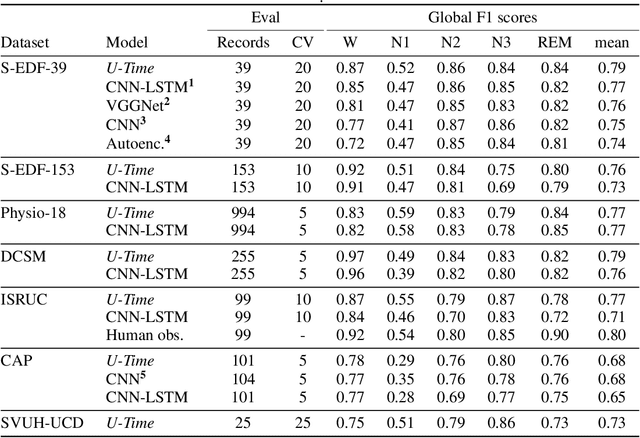
Abstract:Neural networks are becoming more and more popular for the analysis of physiological time-series. The most successful deep learning systems in this domain combine convolutional and recurrent layers to extract useful features to model temporal relations. Unfortunately, these recurrent models are difficult to tune and optimize. In our experience, they often require task-specific modifications, which makes them challenging to use for non-experts. We propose U-Time, a fully feed-forward deep learning approach to physiological time series segmentation developed for the analysis of sleep data. U-Time is a temporal fully convolutional network based on the U-Net architecture that was originally proposed for image segmentation. U-Time maps sequential inputs of arbitrary length to sequences of class labels on a freely chosen temporal scale. This is done by implicitly classifying every individual time-point of the input signal and aggregating these classifications over fixed intervals to form the final predictions. We evaluated U-Time for sleep stage classification on a large collection of sleep electroencephalography (EEG) datasets. In all cases, we found that U-Time reaches or outperforms current state-of-the-art deep learning models while being much more robust in the training process and without requiring architecture or hyperparameter adaptation across tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge