Paschalis Bizopoulos
Comprehensive Comparison of Deep Learning Models for Lung and COVID-19 Lesion Segmentation in CT scans
Sep 10, 2020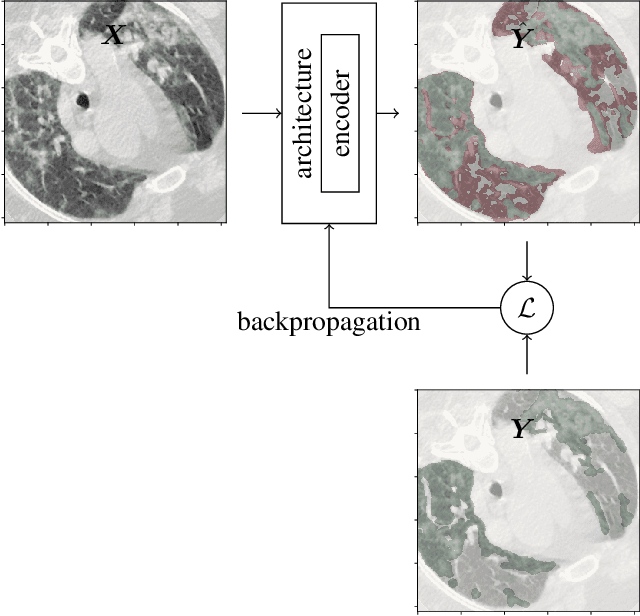
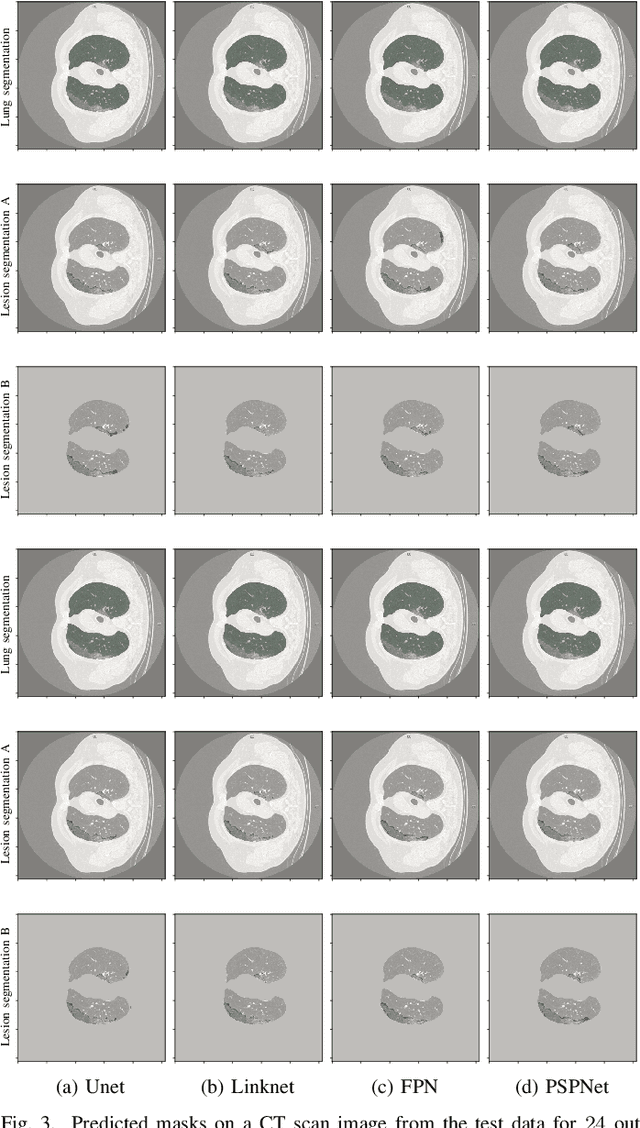



Abstract:Recently there has been an explosion in the use of Deep Learning (DL) methods for medical image segmentation. However the field's reliability is hindered by the lack of a common base of reference for accuracy/performance evaluation and the fact that previous research uses different datasets for evaluation. In this paper, an extensive comparison of DL models for lung and COVID-19 lesion segmentation in Computerized Tomography (CT) scans is presented, which can also be used as a benchmark for testing medical image segmentation models. Four DL architectures (Unet, Linknet, FPN, PSPNet) are combined with 25 randomly initialized and pretrained encoders (variations of VGG, DenseNet, ResNet, ResNext, DPN, MobileNet, Xception, Inception-v4, EfficientNet), to construct 200 tested models. Three experimental setups are conducted for lung segmentation, lesion segmentation and lesion segmentation using the original lung masks. A public COVID-19 dataset with 100 CT scan images (80 for train, 20 for validation) is used for training/validation and a different public dataset consisting of 829 images from 9 CT scan volumes for testing. Multiple findings are provided including the best architecture-encoder models for each experiment as well as mean Dice results for each experiment, architecture and encoder independently. Finally, the upper bounds improvements when using lung masks as a preprocessing step or when using pretrained models are quantified. The source code and 600 pretrained models for the three experiments are provided, suitable for fine-tuning in experimental setups without GPU capabilities.
Sparsely Activated Networks: A new method for decomposing and compressing data
Oct 30, 2019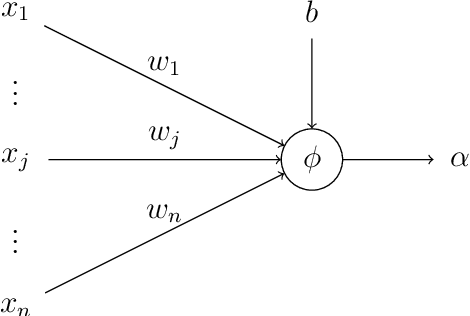


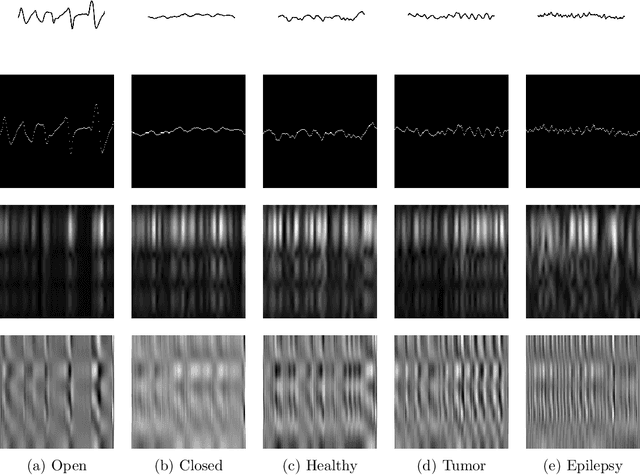
Abstract:Recent literature on unsupervised learning focused on designing structural priors with the aim of learning meaningful features, but without considering the description length of the representations. In this thesis, first we introduce the{\phi}metric that evaluates unsupervised models based on their reconstruction accuracy and the degree of compression of their internal representations. We then present and define two activation functions (Identity, ReLU) as base of reference and three sparse activation functions (top-k absolutes, Extrema-Pool indices, Extrema) as candidate structures that minimize the previously defined metric $\varphi$. We lastly present Sparsely Activated Networks (SANs) that consist of kernels with shared weights that, during encoding, are convolved with the input and then passed through a sparse activation function. During decoding, the same weights are convolved with the sparse activation map and subsequently the partial reconstructions from each weight are summed to reconstruct the input. We compare SANs using the five previously defined activation functions on a variety of datasets (Physionet, UCI-epilepsy, MNIST, FMNIST) and show that models that are selected using $\varphi$ have small description representation length and consist of interpretable kernels.
Sparsely Activated Networks
Jul 12, 2019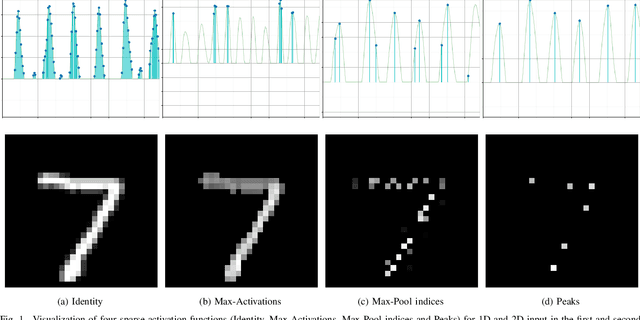
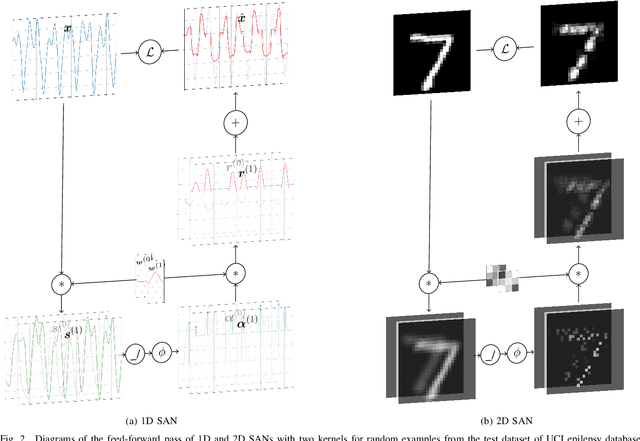
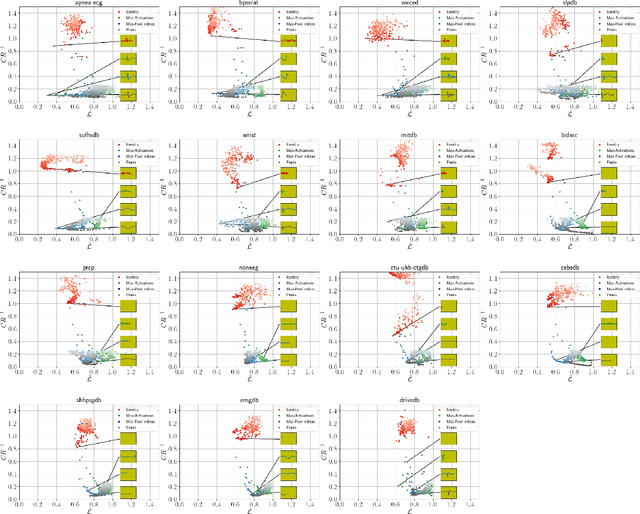
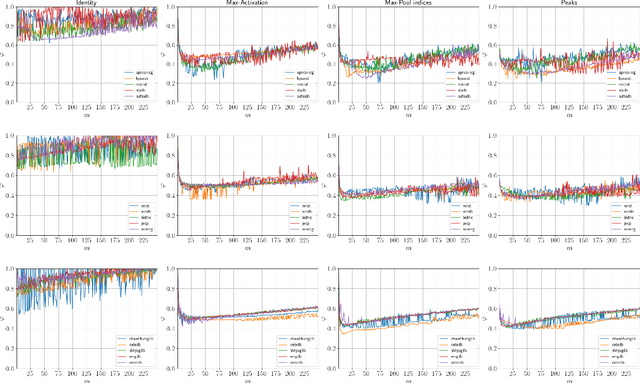
Abstract:Previous literature on unsupervised learning focused on designing structural priors and optimization functions with the aim of learning meaningful features, but without considering the description length of the representations. Here we present Sparsely Activated Networks (SANs), which decompose their input as a sum of sparsely reoccurring patterns of varying amplitude, and combined with a newly proposed metric $\varphi$ they learn representations with minimal description lengths. SANs consist of kernels with shared weights that during encoding are convolved with the input and then passed through a ReLU and a sparse activation function. During decoding, the same weights are convolved with the sparse activation map and the individual reconstructions from each weight are summed to reconstruct the input. We also propose a metric $\varphi$ for model selection that favors models which combine high compression ratio and low reconstruction error and we justify its definition by exploring the hyperparameter space of SANs. We compare four sparse activation functions (Identity, Max-Activations, Max-Pool indices, Peaks) on a variety of datasets and show that SANs learn interpretable kernels that combined with $\varphi$, they minimize the description length of the representations.
Signal2Image Modules in Deep Neural Networks for EEG Classification
May 01, 2019


Abstract:Deep learning has revolutionized computer vision utilizing the increased availability of big data and the power of parallel computational units such as graphical processing units. The vast majority of deep learning research is conducted using images as training data, however the biomedical domain is rich in physiological signals that are used for diagnosis and prediction problems. It is still an open research question how to best utilize signals to train deep neural networks. In this paper we define the term Signal2Image (S2Is) as trainable or non-trainable prefix modules that convert signals, such as Electroencephalography (EEG), to image-like representations making them suitable for training image-based deep neural networks defined as `base models'. We compare the accuracy and time performance of four S2Is (`signal as image', spectrogram, one and two layer Convolutional Neural Networks (CNNs)) combined with a set of `base models' (LeNet, AlexNet, VGGnet, ResNet, DenseNet) along with the depth-wise and 1D variations of the latter. We also provide empirical evidence that the one layer CNN S2I performs better in eleven out of fifteen tested models than non-trainable S2Is for classifying EEG signals and we present visual comparisons of the outputs of the S2Is.
Deep Learning in Cardiology
Feb 22, 2019
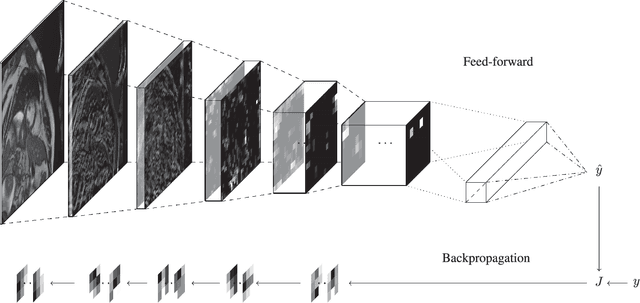


Abstract:The medical field is creating large amount of data that physicians are unable to decipher and use efficiently. Moreover, rule-based expert systems are inefficient in solving complicated medical tasks or for creating insights using big data. Deep learning has emerged as a more accurate and effective technology in a wide range of medical problems such as diagnosis, prediction and intervention. Deep learning is a representation learning method that consists of layers that transform the data non-linearly, thus, revealing hierarchical relationships and structures. In this review we survey deep learning application papers that use structured data, signal and imaging modalities from cardiology. We discuss the advantages and limitations of applying deep learning in cardiology that also apply in medicine in general, while proposing certain directions as the most viable for clinical use.
* 27 pages, 2 figures, 10 tables
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge