Natalie Gangai
A Novel Patch-Based TDA Approach for Computed Tomography
Dec 13, 2025



Abstract:The development of machine learning (ML) models based on computed tomography (CT) imaging modality has been a major focus of recent research in the medical imaging domain. Incorporating robust feature engineering approach can highly improve the performance of these models. Topological data analysis (TDA), a recent development based on the mathematical field of algebraic topology, mainly focuses on the data from a topological perspective, extracting deeper insight and higher dimensional structures from the data. Persistent homology (PH), a fundamental tool in the area of TDA, can extract topological features such as connected components, cycles and voids from the data. A popular approach to construct PH from 3D CT images is to utilize the 3D cubical complex filtration, a method adapted for grid-structured data. However, this approach may not always yield the best performance and can suffer from computational complexity with higher resolution CT images. This study introduces a novel patch-based PH construction approach tailored for volumetric medical imaging data, in particular CT modality. A wide range of experiments has been conducted on several datasets of 3D CT images to comprehensively analyze the performance of the proposed method with various parameters and benchmark it against the 3D cubical complex algorithm. Our results highlight the dominance of the patch-based TDA approach in terms of both classification performance and time-efficiency. The proposed approach outperformed the cubical complex method, achieving average improvement of 10.38%, 6.94%, 2.06%, 11.58%, and 8.51% in accuracy, AUC, sensitivity, specificity, and F1 score, respectively, across all datasets. Finally, we provide a convenient python package, Patch-TDA, to facilitate the utilization of the proposed approach.
Finding Reproducible and Prognostic Radiomic Features in Variable Slice Thickness Contrast Enhanced CT of Colorectal Liver Metastases
Jan 20, 2025



Abstract:Establishing the reproducibility of radiomic signatures is a critical step in the path to clinical adoption of quantitative imaging biomarkers; however, radiomic signatures must also be meaningfully related to an outcome of clinical importance to be of value for personalized medicine. In this study, we analyze both the reproducibility and prognostic value of radiomic features extracted from the liver parenchyma and largest liver metastases in contrast enhanced CT scans of patients with colorectal liver metastases (CRLM). A prospective cohort of 81 patients from two major US cancer centers was used to establish the reproducibility of radiomic features extracted from images reconstructed with different slice thicknesses. A publicly available, single-center cohort of 197 preoperative scans from patients who underwent hepatic resection for treatment of CRLM was used to evaluate the prognostic value of features and models to predict overall survival. A standard set of 93 features was extracted from all images, with a set of eight different extractor settings. The feature extraction settings producing the most reproducible, as well as the most prognostically discriminative feature values were highly dependent on both the region of interest and the specific feature in question. While the best overall predictive model was produced using features extracted with a particular setting, without accounting for reproducibility, (C-index = 0.630 (0.603--0.649)) an equivalent-performing model (C-index = 0.629 (0.605--0.645)) was produced by pooling features from all extraction settings, and thresholding features with low reproducibility ($\mathrm{CCC} \geq 0.85$), prior to feature selection. Our findings support a data-driven approach to feature extraction and selection, preferring the inclusion of many features, and narrowing feature selection based on reproducibility when relevant data is available.
* Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://melba-journal.org/2024:032
Transformer-based segmentation of adnexal lesions and ovarian implants in CT images
Jun 25, 2024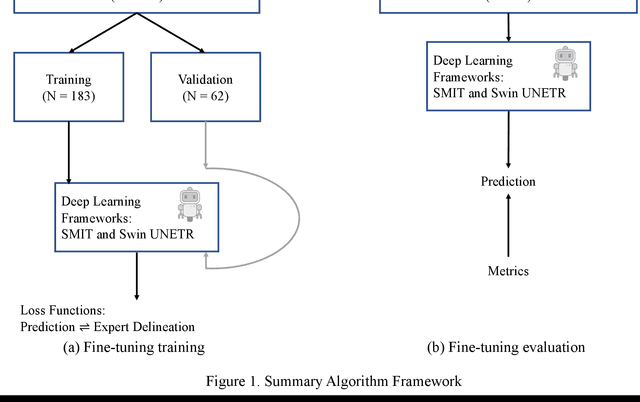
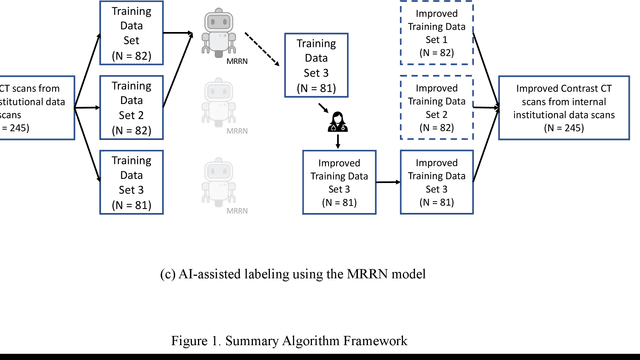
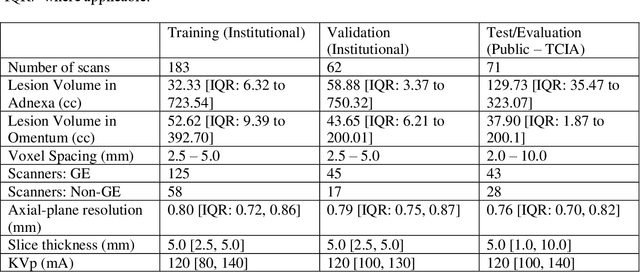
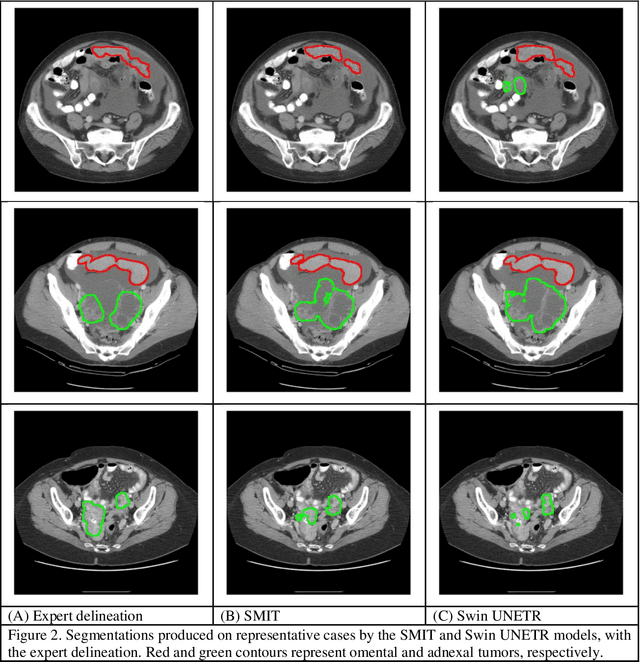
Abstract:Two self-supervised pretrained transformer-based segmentation models (SMIT and Swin UNETR) fine-tuned on a dataset of ovarian cancer CT images provided reasonably accurate delineations of the tumors in an independent test dataset. Tumors in the adnexa were segmented more accurately by both transformers (SMIT and Swin UNETR) than the omental implants. AI-assisted labeling performed on 72 out of 245 omental implants resulted in smaller manual editing effort of 39.55 mm compared to full manual correction of partial labels of 106.49 mm and resulted in overall improved accuracy performance. Both SMIT and Swin UNETR did not generate any false detection of omental metastases in the urinary bladder and relatively few false detections in the small bowel, with 2.16 cc on average for SMIT and 7.37 cc for Swin UNETR respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge