Milda Pocevičiūtė
Detecting Domain Shift in Multiple Instance Learning for Digital Pathology Using Fréchet Domain Distance
May 16, 2024Abstract:Multiple-instance learning (MIL) is an attractive approach for digital pathology applications as it reduces the costs related to data collection and labelling. However, it is not clear how sensitive MIL is to clinically realistic domain shifts, i.e., differences in data distribution that could negatively affect performance, and if already existing metrics for detecting domain shifts work well with these algorithms. We trained an attention-based MIL algorithm to classify whether a whole-slide image of a lymph node contains breast tumour metastases. The algorithm was evaluated on data from a hospital in a different country and various subsets of this data that correspond to different levels of domain shift. Our contributions include showing that MIL for digital pathology is affected by clinically realistic differences in data, evaluating which features from a MIL model are most suitable for detecting changes in performance, and proposing an unsupervised metric named Fr\'echet Domain Distance (FDD) for quantification of domain shifts. Shift measure performance was evaluated through the mean Pearson correlation to change in classification performance, where FDD achieved 0.70 on 10-fold cross-validation models. The baselines included Deep ensemble, Difference of Confidence, and Representation shift which resulted in 0.45, -0.29, and 0.56 mean Pearson correlation, respectively. FDD could be a valuable tool for care providers and vendors who need to verify if a MIL system is likely to perform reliably when implemented at a new site, without requiring any additional annotations from pathologists.
Can uncertainty boost the reliability of AI-based diagnostic methods in digital pathology?
Dec 17, 2021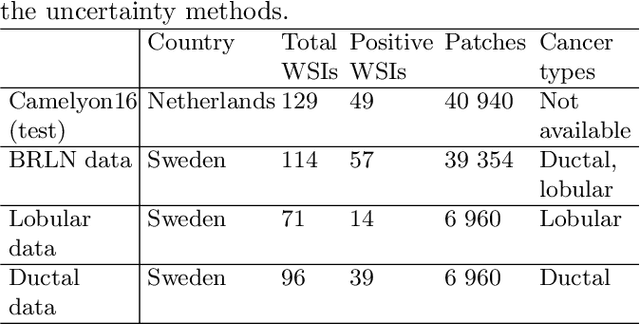


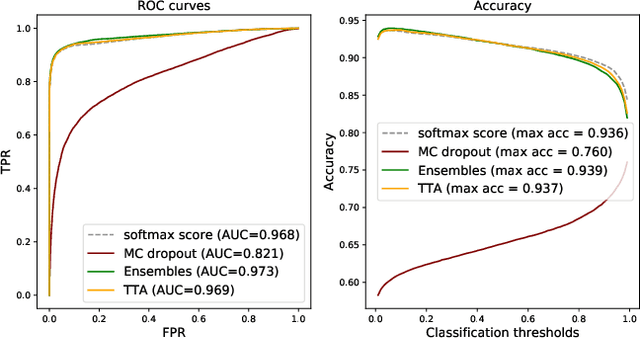
Abstract:Deep learning (DL) has shown great potential in digital pathology applications. The robustness of a diagnostic DL-based solution is essential for safe clinical deployment. In this work we evaluate if adding uncertainty estimates for DL predictions in digital pathology could result in increased value for the clinical applications, by boosting the general predictive performance or by detecting mispredictions. We compare the effectiveness of model-integrated methods (MC dropout and Deep ensembles) with a model-agnostic approach (Test time augmentation, TTA). Moreover, four uncertainty metrics are compared. Our experiments focus on two domain shift scenarios: a shift to a different medical center and to an underrepresented subtype of cancer. Our results show that uncertainty estimates can add some reliability and reduce sensitivity to classification threshold selection. While advanced metrics and deep ensembles perform best in our comparison, the added value over simpler metrics and TTA is small. Importantly, the benefit of all evaluated uncertainty estimation methods is diminished by domain shift.
Unsupervised anomaly detection in digital pathology using GANs
Mar 16, 2021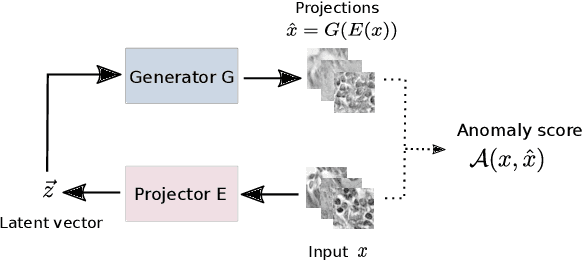
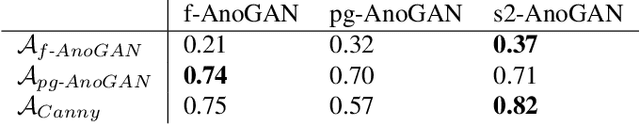
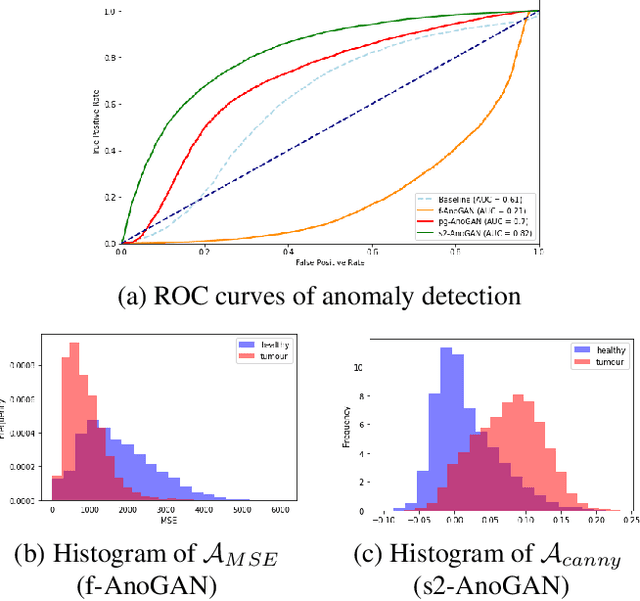

Abstract:Machine learning (ML) algorithms are optimized for the distribution represented by the training data. For outlier data, they often deliver predictions with equal confidence, even though these should not be trusted. In order to deploy ML-based digital pathology solutions in clinical practice, effective methods for detecting anomalous data are crucial to avoid incorrect decisions in the outlier scenario. We propose a new unsupervised learning approach for anomaly detection in histopathology data based on generative adversarial networks (GANs). Compared to the existing GAN-based methods that have been used in medical imaging, the proposed approach improves significantly on performance for pathology data. Our results indicate that histopathology imagery is substantially more complex than the data targeted by the previous methods. This complexity requires not only a more advanced GAN architecture but also an appropriate anomaly metric to capture the quality of the reconstructed images.
Survey of XAI in digital pathology
Aug 14, 2020
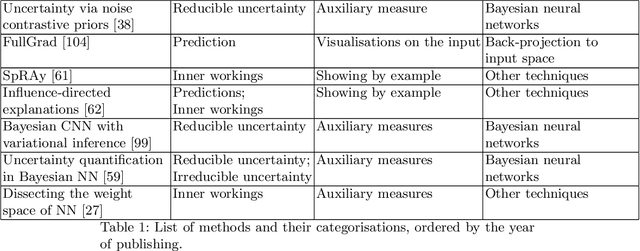
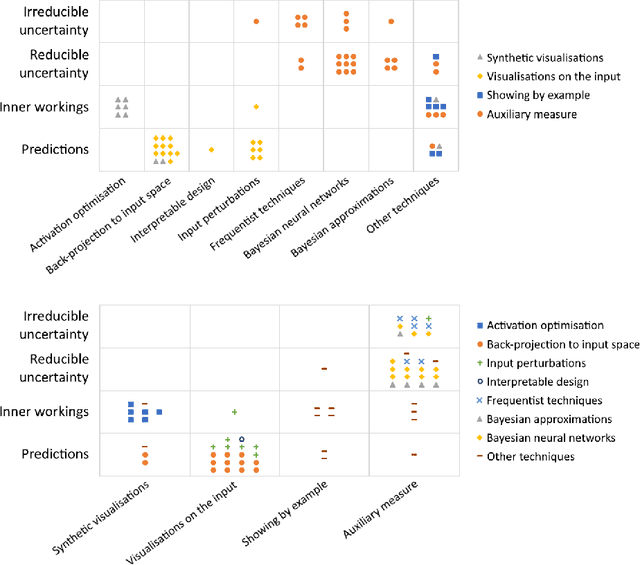
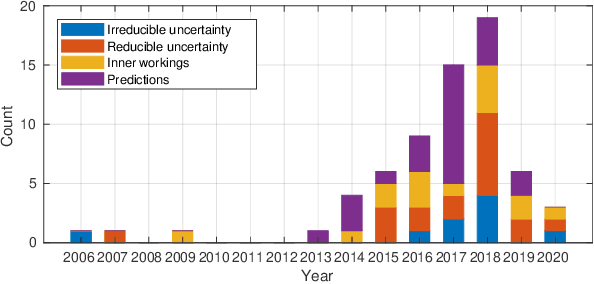
Abstract:Artificial intelligence (AI) has shown great promise for diagnostic imaging assessments. However, the application of AI to support medical diagnostics in clinical routine comes with many challenges. The algorithms should have high prediction accuracy but also be transparent, understandable and reliable. Thus, explainable artificial intelligence (XAI) is highly relevant for this domain. We present a survey on XAI within digital pathology, a medical imaging sub-discipline with particular characteristics and needs. The review includes several contributions. Firstly, we give a thorough overview of current XAI techniques of potential relevance for deep learning methods in pathology imaging, and categorise them from three different aspects. In doing so, we incorporate uncertainty estimation methods as an integral part of the XAI landscape. We also connect the technical methods to the specific prerequisites in digital pathology and present findings to guide future research efforts. The survey is intended for both technical researchers and medical professionals, one of the objectives being to establish a common ground for cross-disciplinary discussions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge