Michal Španěl
From Synthetic Data to Real Restorations: Diffusion Model for Patient-specific Dental Crown Completion
Mar 27, 2026Abstract:We present ToothCraft, a diffusion-based model for the contextual generation of tooth crowns, trained on artificially created incomplete teeth. Building upon recent advancements in conditioned diffusion models for 3D shapes, we developed a model capable of an automated tooth crown completion conditioned on local anatomical context. To address the lack of training data for this task, we designed an augmentation pipeline that generates incomplete tooth geometries from a publicly available dataset of complete dental arches (3DS, ODD). By synthesising a diverse set of training examples, our approach enables robust learning across a wide spectrum of tooth defects. Experimental results demonstrate the strong capability of our model to reconstruct complete tooth crowns, achieving an intersection over union (IoU) of 81.8% and a Chamfer Distance (CD) of 0.00034 on synthetically damaged testing restorations. Our experiments demonstrate that the model can be applied directly to real-world cases, effectively filling in incomplete teeth, while generated crowns show minimal intersection with the opposing dentition, thus reducing the risk of occlusal interference. Access to the code, model weights, and dataset information will be available at: https://github.com/ikarus1211/VISAPP_ToothCraft
Segmentation of Defective Skulls from CT Data for Tissue Modelling
Nov 20, 2019
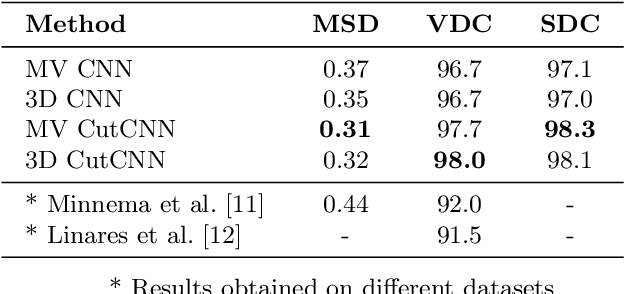


Abstract:In this work we present a method of automatic segmentation of defective skulls for custom cranial implant design and 3D printing purposes. Since such tissue models are usually required in patient cases with complex anatomical defects and variety of external objects present in the acquired data, most deep learning-based approaches fall short because it is not possible to create a sufficient training dataset that would encompass the spectrum of all possible structures. Because CNN segmentation experiments in this application domain have been so far limited to simple patch-based CNN architectures, we first show how the usage of the encoder-decoder architecture can substantially improve the segmentation accuracy. Then, we show how the number of segmentation artifacts, which usually require manual corrections, can be further reduced by adding a boundary term to CNN training and by globally optimizing the segmentation with graph-cut. Finally, we show that using the proposed method, 3D segmentation accurate enough for clinical application can be achieved with 2D CNN architectures as well as their 3D counterparts.
Segmentation of Head and Neck Organs at Risk Using CNN with Batch Dice Loss
Dec 06, 2018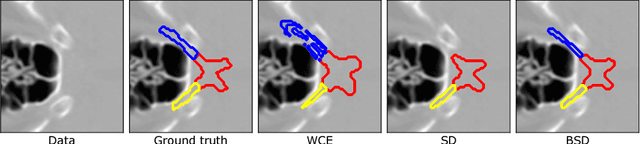
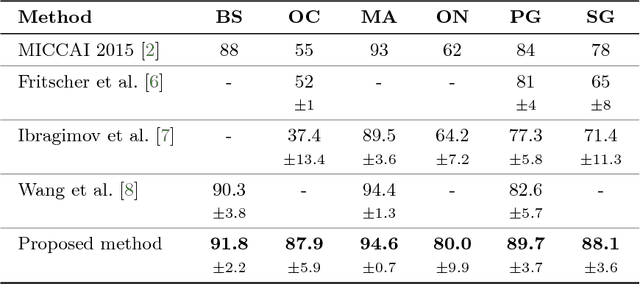
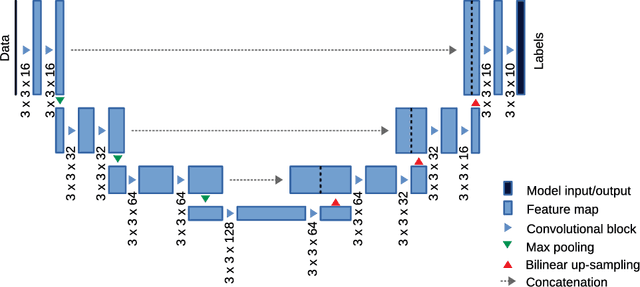
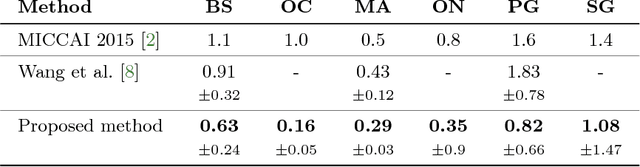
Abstract:This paper deals with segmentation of organs at risk (OAR) in head and neck area in CT images which is a crucial step for reliable intensity modulated radiotherapy treatment. We introduce a convolution neural network with encoder-decoder architecture and a new loss function, the batch soft Dice loss function, used to train the network. The resulting model produces segmentations of every OAR in the public MICCAI 2015 Head And Neck Auto-Segmentation Challenge dataset. Despite the heavy class imbalance in the data, we improve accuracy of current state-of-the-art methods by 0.33 mm in terms of average surface distance and by 0.11 in terms of Dice overlap coefficient on average.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge