Michael Doukas
EPIC-Survival: End-to-end Part Inferred Clustering for Survival Analysis, Featuring Prognostic Stratification Boosting
Jan 28, 2021
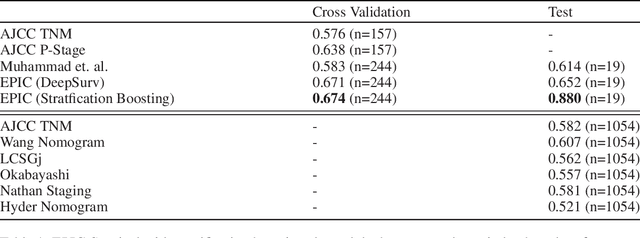
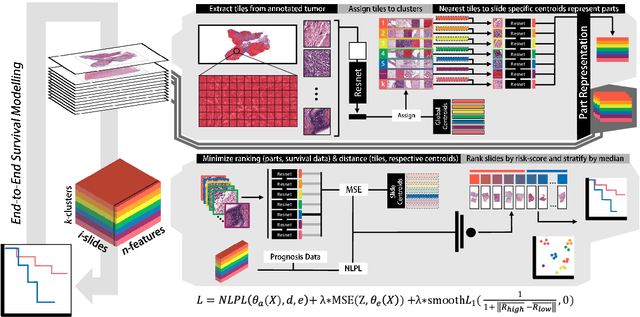
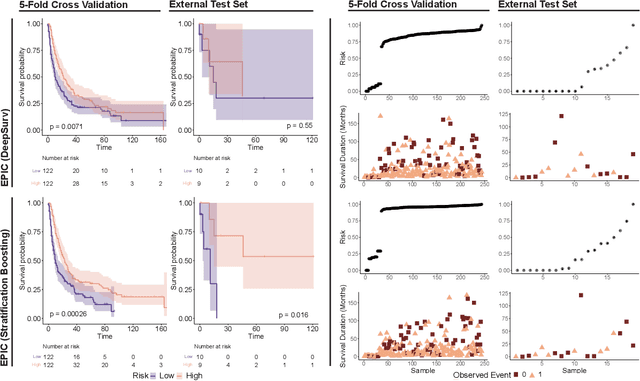
Abstract:Histopathology-based survival modelling has two major hurdles. Firstly, a well-performing survival model has minimal clinical application if it does not contribute to the stratification of a cancer patient cohort into different risk groups, preferably driven by histologic morphologies. In the clinical setting, individuals are not given specific prognostic predictions, but are rather predicted to lie within a risk group which has a general survival trend. Thus, It is imperative that a survival model produces well-stratified risk groups. Secondly, until now, survival modelling was done in a two-stage approach (encoding and aggregation). The massive amount of pixels in digitized whole slide images were never utilized to their fullest extent due to technological constraints on data processing, forcing decoupled learning. EPIC-Survival bridges encoding and aggregation into an end-to-end survival modelling approach, while introducing stratification boosting to encourage the model to not only optimize ranking, but also to discriminate between risk groups. In this study we show that EPIC-Survival performs better than other approaches in modelling intrahepatic cholangiocarcinoma, a historically difficult cancer to model. Further, we show that stratification boosting improves further improves model performance, resulting in a concordance-index of 0.880 on a held-out test set. Finally, we were able to identify specific histologic differences, not commonly sought out in ICC, between low and high risk groups.
Towards Unsupervised Cancer Subtyping: Predicting Prognosis Using A Histologic Visual Dictionary
Mar 12, 2019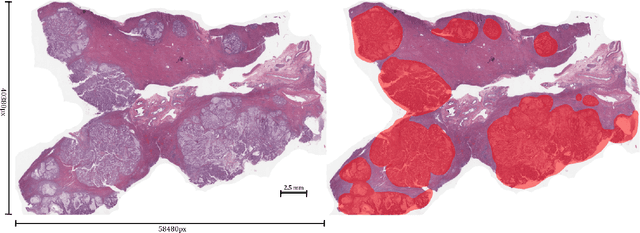
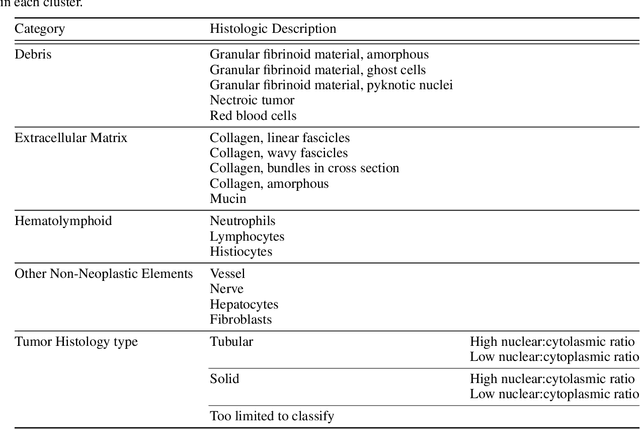
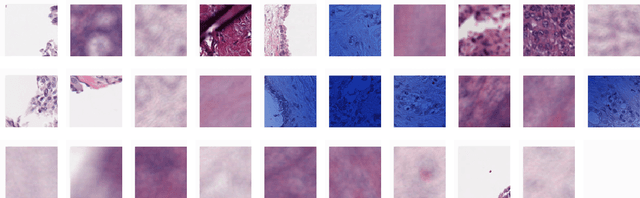
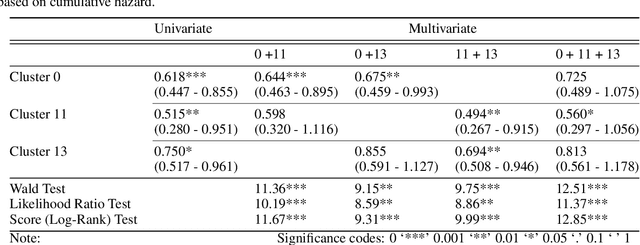
Abstract:Unlike common cancers, such as those of the prostate and breast, tumor grading in rare cancers is difficult and largely undefined because of small sample sizes, the sheer volume of time needed to undertake on such a task, and the inherent difficulty of extracting human-observed patterns. One of the most challenging examples is intrahepatic cholangiocarcinoma (ICC), a primary liver cancer arising from the biliary system, for which there is well-recognized tumor heterogeneity and no grading paradigm or prognostic biomarkers. In this paper, we propose a new unsupervised deep convolutional autoencoder-based clustering model that groups together cellular and structural morphologies of tumor in 246 ICC digitized whole slides, based on visual similarity. From this visual dictionary of histologic patterns, we use the clusters as covariates to train Cox-proportional hazard survival models. In univariate analysis, three clusters were significantly associated with recurrence-free survival. Combinations of these clusters were significant in multivariate analysis. In a multivariate analysis of all clusters, five showed significance to recurrence-free survival, however the overall model was not measured to be significant. Finally, a pathologist assigned clinical terminology to the significant clusters in the visual dictionary and found evidence supporting the hypothesis that collagen-enriched fibrosis plays a role in disease severity. These results offer insight into the future of cancer subtyping and show that computational pathology can contribute to disease prognostication, especially in rare cancers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge